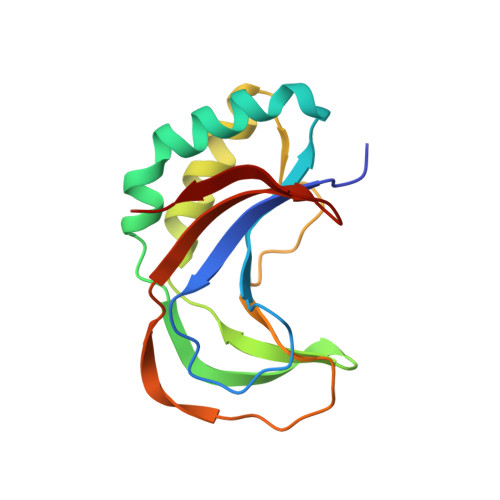
Crystal Structure of the C-Terminal 2',5'-Phosphodiesterase Domain of Group a Rotavirus Protein Vp3.
Brandmann, T., Jinek, M.(2015) Proteins 83: 997
- PubMed: 25758703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24794
- Primary Citation Related Structures:
5AF2 - PubMed Abstract:
In response to viral infections, the mammalian innate immune system induces the production of the second messenger 2'-5' oligoadenylate (2-5A) to activate latent ribonuclease L (RNase L) that restricts viral replication and promotes apoptosis. A subset of rotaviruses and coronaviruses encode 2',5'-phosphodiesterase enzymes that hydrolyze 2-5A, thereby inhibiting RNase L activation. We report the crystal structure of the 2',5'-phosphodiesterase domain of group A rotavirus protein VP3 at 1.39 Å resolution. The structure exhibits a 2H phosphoesterase fold and reveals conserved active site residues, providing insights into the mechanism of 2-5A degradation in viral evasion of host innate immunity.
- Department of Biochemistry, University of Zurich, Zurich, Switzerland.
Organizational Affiliation: