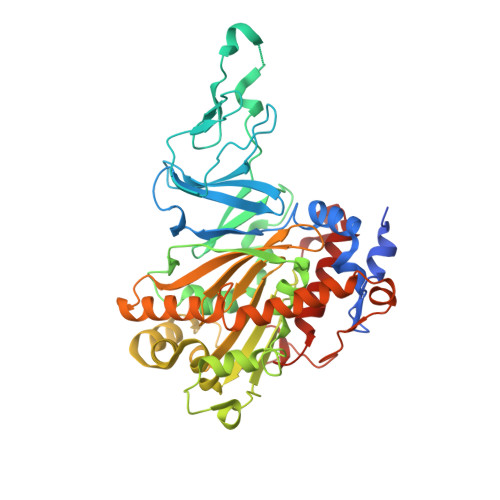
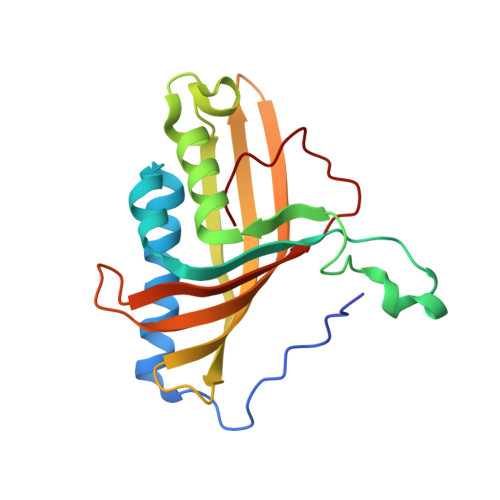
Structural Basis of the Enhanced Pollutant-Degrading Capabilities of an Engineered Biphenyl Dioxygenase.
Dhindwal, S., Gomez-Gil, L., Neau, D.B., Pham, T.T.M., Sylvestre, M., Eltis, L.D., Bolin, J.T., Kumar, P.(2016) J Bacteriol 198: 1499
- PubMed: 26953337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00952-15
- Primary Citation Related Structures:
5AEU, 5AEW - PubMed Abstract:
Biphenyl dioxygenase, the first enzyme of the biphenyl catabolic pathway, is a major determinant of which polychlorinated biphenyl (PCB) congeners are metabolized by a given bacterial strain. Ongoing efforts aim to engineer BphAE, the oxygenase component of the enzyme, to efficiently transform a wider range of congeners. BphAEII9, a variant of BphAELB400 in which a seven-residue segment, (335)TFNNIRI(341), has been replaced by the corresponding segment of BphAEB356, (333)GINTIRT(339), transforms a broader range of PCB congeners than does either BphAELB400 or BphAEB356, including 2,6-dichlorobiphenyl, 3,3'-dichlorobiphenyl, 4,4'-dichlorobiphenyl, and 2,3,4'-trichlorobiphenyl. To understand the structural basis of the enhanced activity of BphAEII9, we have determined the three-dimensional structure of this variant in substrate-free and biphenyl-bound forms. Structural comparison with BphAELB400 reveals a flexible active-site mouth and a relaxed substrate binding pocket in BphAEII9 that allow it to bind different congeners and which could be responsible for the enzyme's altered specificity. Biochemical experiments revealed that BphAEII9 transformed 2,3,4'-trichlorobiphenyl and 2,2',5,5'-tetrachlorobiphenyl more efficiently than did BphAELB400 and BphAEB356 BphAEII9 also transformed the insecticide dichlorodiphenyltrichloroethane (DDT) more efficiently than did either parental enzyme (apparent kcat/Km of 2.2 ± 0.5 mM(-1) s(-1), versus 0.9 ± 0.5 mM(-1) s(-1) for BphAEB356). Studies of docking of the enzymes with these three substrates provide insight into the structural basis of the different substrate selectivities and regiospecificities of the enzymes. Biphenyl dioxygenase is the first enzyme of the biphenyl degradation pathway that is involved in the degradation of polychlorinated biphenyls. Attempts have been made to identify the residues that influence the enzyme activity for the range of substrates among various species. In this study, we have done a structural study of one variant of this enzyme that was produced by family shuffling of genes from two different species. Comparison of the structure of this variant with those of the parent enzymes provided an important insight into the molecular basis for the broader substrate preference of this enzyme. The structural and functional details gained in this study can be utilized to further engineer desired enzymatic activity, producing more potent enzymes.
- Department of Biotechnology, Indian Institute of Technology Roorkee, Roorkee, Uttarakhand, India.
Organizational Affiliation: