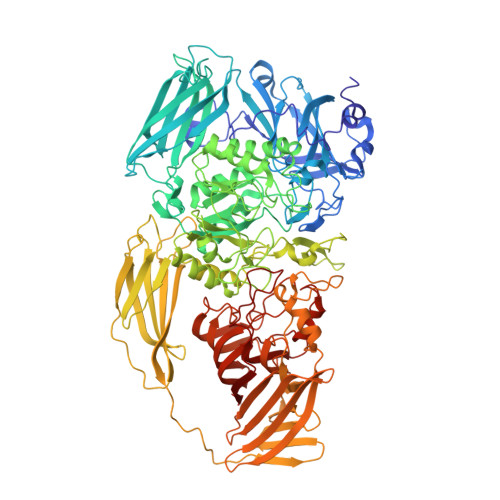
2.2 A Resolution Cryo-Em Structure of Beta-Galactosidase in Complex with a Cell-Permeant Inhibitor
Bartesaghi, A., Merk, A., Banerjee, S., Matthies, D., Wu, X., Milne, J., Subramaniam, S.(2015) Science 348: 1147
- PubMed: 25953817 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aab1576
- Primary Citation Related Structures:
5A1A - PubMed Abstract:
Cryo-electron microscopy (cryo-EM) is rapidly emerging as a powerful tool for protein structure determination at high resolution. Here we report the structure of a complex between Escherichia coli β-galactosidase and the cell-permeant inhibitor phenylethyl β-D-thiogalactopyranoside (PETG), determined by cryo-EM at an average resolution of ~2.2 angstroms (Å). Besides the PETG ligand, we identified densities in the map for ~800 water molecules and for magnesium and sodium ions. Although it is likely that continued advances in detector technology may further enhance resolution, our findings demonstrate that preparation of specimens of adequate quality and intrinsic protein flexibility, rather than imaging or image-processing technologies, now represent the major bottlenecks to routinely achieving resolutions close to 2 Å using single-particle cryo-EM.
- Laboratory of Cell Biology, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: