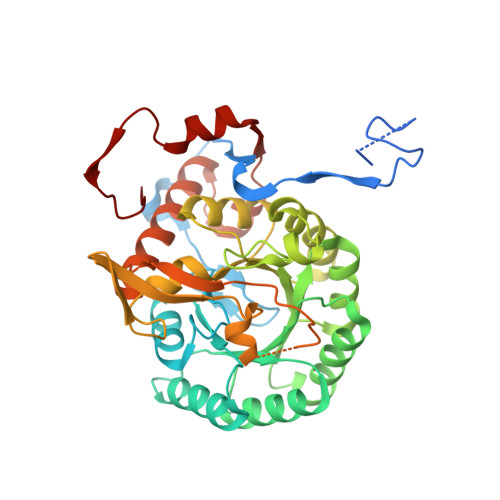
Mycobacterium tuberculosis IMPDH in Complexes with Substrates, Products and Antitubercular Compounds.
Makowska-Grzyska, M., Kim, Y., Gorla, S.K., Wei, Y., Mandapati, K., Zhang, M., Maltseva, N., Modi, G., Boshoff, H.I., Gu, M., Aldrich, C., Cuny, G.D., Hedstrom, L., Joachimiak, A.(2015) PLoS One 10: e0138976-e0138976
- PubMed: 26440283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0138976
- Primary Citation Related Structures:
4ZQM, 4ZQN, 4ZQO, 4ZQP, 4ZQR - PubMed Abstract:
Tuberculosis (TB) remains a worldwide problem and the need for new drugs is increasingly more urgent with the emergence of multidrug- and extensively-drug resistant TB. Inosine 5'-monophosphate dehydrogenase 2 (IMPDH2) from Mycobacterium tuberculosis (Mtb) is an attractive drug target. The enzyme catalyzes the conversion of inosine 5'-monophosphate into xanthosine 5'-monophosphate with the concomitant reduction of NAD+ to NADH. This reaction controls flux into the guanine nucleotide pool. We report seventeen selective IMPDH inhibitors with antitubercular activity. The crystal structures of a deletion mutant of MtbIMPDH2 in the apo form and in complex with the product XMP and substrate NAD+ are determined. We also report the structures of complexes with IMP and three structurally distinct inhibitors, including two with antitubercular activity. These structures will greatly facilitate the development of MtbIMPDH2-targeted antibiotics.
- Center for Structural Genomics of Infectious Diseases, University of Chicago, Chicago, IL, United States of America.
Organizational Affiliation: