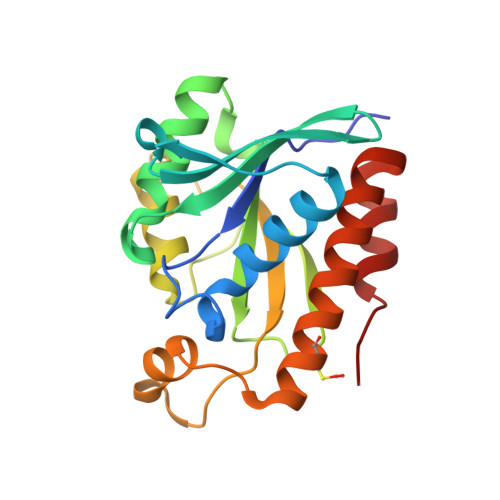
Unraveling the stereochemical and dynamic aspects of the catalytic site of bacterial peptidyl-tRNA hydrolase.
Kabra, A., Shahid, S., Pal, R.K., Yadav, R., Pulavarti, S.V., Jain, A., Tripathi, S., Arora, A.(2017) RNA 23: 202-216
- PubMed: 28096445 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.057620.116
- Primary Citation Related Structures:
4Z86, 4ZXP, 5B6J, 5IKE, 5IMB - PubMed Abstract:
Bacterial peptidyl-tRNA hydrolase (Pth; EC 3.1.1.29) hydrolyzes the peptidyl-tRNAs accumulated in the cytoplasm and thereby prevents cell death by alleviating tRNA starvation. X-ray and NMR studies of Vibrio cholerae Pth (VcPth) and mutants of its key residues involved in catalysis show that the activity and selectivity of the protein depends on the stereochemistry and dynamics of residues H24, D97, N118, and N14. D97-H24 interaction is critical for activity because it increases the nucleophilicity of H24. The N118 and N14 have orthogonally competing interactions with H24, both of which reduce the nucleophilicity of H24 and are likely to be offset by positioning of a peptidyl-tRNA substrate. The region proximal to H24 and the lid region exhibit slow motions that may assist in accommodating the substrate. Helix α3 exhibits a slow wobble with intermediate time scale motions of its N-cap residue N118, which may work as a flypaper to position the scissile ester bond of the substrate. Overall, the dynamics of interactions between the side chains of N14, H24, D97, and N118, control the catalysis of substrate by this enzyme.
- Molecular and Structural Biology Division, CSIR-Central Drug Research Institute, Lucknow 226031, India.
Organizational Affiliation: