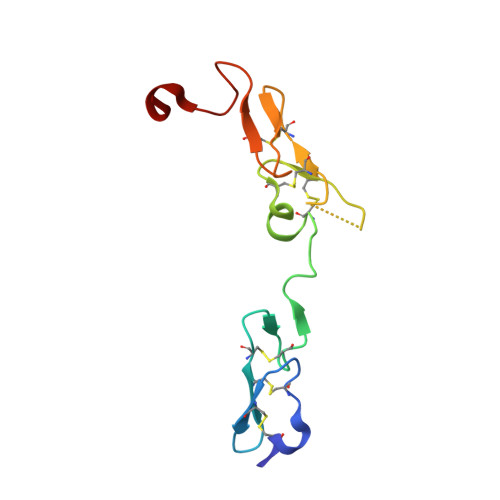
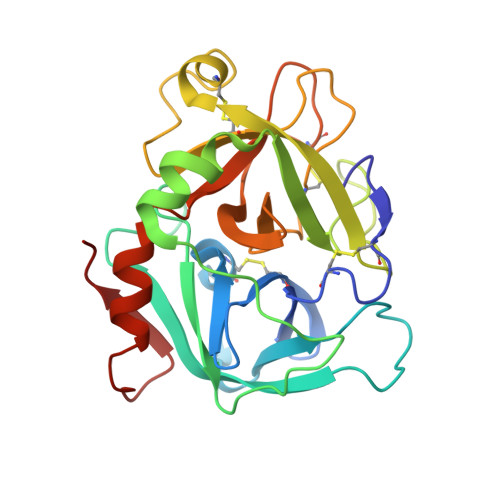
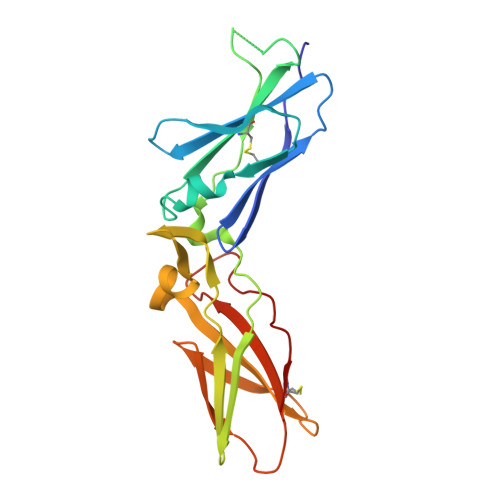
Molecular Basis of Enhanced Activity in Factor VIIa-Trypsin Variants Conveys Insights into Tissue Factor-mediated Allosteric Regulation of Factor VIIa Activity.
Sorensen, A.B., Madsen, J.J., Svensson, L.A., Pedersen, A.A., stergaard, H., Overgaard, M.T., Olsen, O.H., Gandhi, P.S.(2016) J Biological Chem 291: 4671-4683
- PubMed: 26694616 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.698613
- Primary Citation Related Structures:
4YLQ, 4Z6A, 4ZMA - PubMed Abstract:
The complex of coagulation factor VIIa (FVIIa), a trypsin-like serine protease, and membrane-bound tissue factor (TF) initiates blood coagulation upon vascular injury. Binding of TF to FVIIa promotes allosteric conformational changes in the FVIIa protease domain and improves its catalytic properties. Extensive studies have revealed two putative pathways for this allosteric communication. Here we provide further details of this allosteric communication by investigating FVIIa loop swap variants containing the 170 loop of trypsin that display TF-independent enhanced activity. Using x-ray crystallography, we show that the introduced 170 loop from trypsin directly interacts with the FVIIa active site, stabilizing segment 215-217 and activation loop 3, leading to enhanced activity. Molecular dynamics simulations and novel fluorescence quenching studies support that segment 215-217 conformation is pivotal to the enhanced activity of the FVIIa variants. We speculate that the allosteric regulation of FVIIa activity by TF binding follows a similar path in conjunction with protease domain N terminus insertion, suggesting a more complete molecular basis of TF-mediated allosteric enhancement of FVIIa activity.
- From Global Research, Novo Nordisk A/S, 2760 Måløv, Denmark, Department of Chemistry and Bioscience, Aalborg University, 9220 Aalborg, Denmark, and.
Organizational Affiliation: