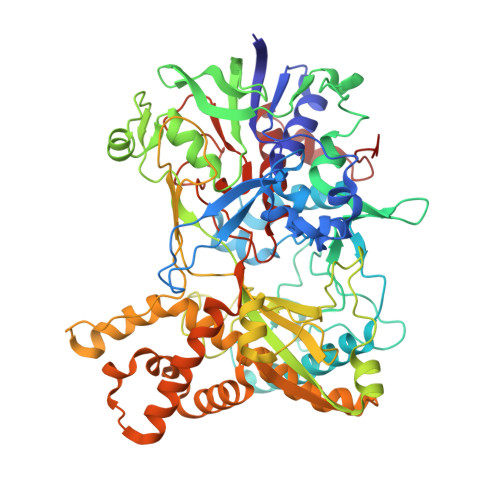
A Mimivirus Enzyme that Participates in Viral Entry.
Klose, T., Herbst, D.A., Zhu, H., Max, J.P., Kenttamaa, H.I., Rossmann, M.G.(2015) Structure 23: 1058-1065
- PubMed: 25982526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.03.023
- Primary Citation Related Structures:
4Z24, 4Z25, 4Z26 - PubMed Abstract:
Mimivirus was initially identified as a bacterium because its dense, 125-nm-long fibers stained Gram-positively. These fibers probably play a role during the infection of some host cells. The normal hosts of Mimivirus are unknown, but in the laboratory Mimivirus is usually propagated in amoeba. The structure of R135, a major component of the fibrous outer layer of Mimivirus, has been determined to 2-Å resolution. The protein's structure is similar to that of members of the glucose-methanol-choline oxidoreductase family, which have an N-terminal FAD binding domain and a C-terminal substrate recognition domain. The closest homolog to R135 is an aryl-alcohol oxidase that participates in lignin biodegradation of plant cell walls. Thus R135 might participate in the degradation of their normal hosts, including some lignin-containing algae.
- Department of Biological Sciences, Purdue University, 240 South Martin Jischke Drive, West Lafayette, IN 47907-2032, USA.
Organizational Affiliation: