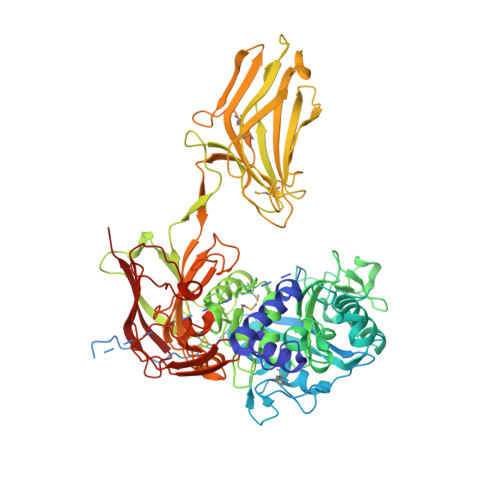
Structural Basis for Latency and Function of Immune Inhibitor A Metallopeptidase, a Modulator of the Bacillus anthracis Secretome.
Arolas, J.L., Goulas, T., Pomerantsev, A.P., Leppla, S.H., Gomis-Ruth, F.X.(2016) Structure 24: 25-36
- PubMed: 26745529 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.10.015
- Primary Citation Related Structures:
4YU5, 4YU6 - PubMed Abstract:
Immune inhibitor A(InhA)-type metallopeptidases are potential virulence factors secreted by members of the Bacillus cereus group. Two paralogs from anthrax-causing Bacillus anthracis (BaInhA1 and BaInhA2) were shown to degrade host tissue proteins with broad substrate specificity. Analysis of their activation mechanism and the crystal structure of a zymogenic BaInhA2 variant revealed a ∼750-residue four-domain structure featuring a pro-peptide, a catalytic domain, a domain reminiscent of viral envelope glycoproteins, and a MAM domain grafted into the latter. This domain, previously found only in eukaryotes, is required for proper protein expression in B. anthracis and evinces certain flexibility. Latency is uniquely modulated by the N-terminal segment of the pro-peptide, which binds the catalytic zinc through its α-amino group and occupies the primed side of the active-site cleft. The present results further our understanding of the modus operandi of an anthrax secretome regulator.
- Proteolysis Lab, Department of Structural Biology ("María de Maeztu" Unit of Excellence), Molecular Biology Institute of Barcelona, Spanish Research Council (CSIC), Barcelona Science Park, Helix Building, Baldiri Reixac, 15-21, 08028 Barcelona, Spain.
Organizational Affiliation: