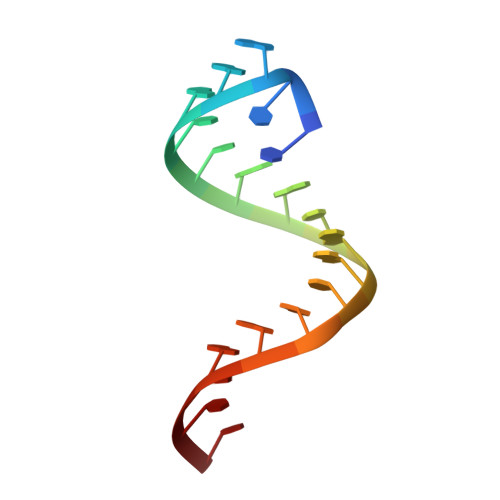
Structural Insights Reveal the Dynamics of the Repeating r(CAG) Transcript Found in Huntington's Disease (HD) and Spinocerebellar Ataxias (SCAs)
Tawani, A., Kumar, A.(2015) PLoS One 10: e0131788-e0131788
- PubMed: 26148061 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0131788
- Primary Citation Related Structures:
2MS5, 4YN6 - PubMed Abstract:
In humans, neurodegenerative disorders such as Huntington's disease (HD) and many spinocerebellar ataxias (SCAs) have been found to be associated with CAG trinucleotide repeat expansion. An important RNA-mediated mechanism that causes these diseases involves the binding of the splicing regulator protein MBNL1 (Muscleblind-like 1 protein) to expanded r(CAG) repeats. Moreover, mutant huntingtin protein translated from expanded r(CAG) also yields toxic effects. To discern the role of mutant RNA in these diseases, it is essential to gather information about its structure. Detailed insight into the different structures and conformations adopted by these mutant transcripts is vital for developing therapeutics targeting them. Here, we report the crystal structure of an RNA model with a r(CAG) motif, which is complemented by an NMR-based solution structure obtained from restrained Molecular Dynamics (rMD) simulation studies. Crystal structure data of the RNA model resolved at 2.3 Å reveals non-canonical pairing of adenine in 5´-CAG/3´-GAC motif samples in different syn and anti conformations. The overall RNA structure has helical parameters intermediate to the A- and B-forms of nucleic acids due to the global widening of major grooves and base-pair preferences near internal AA loops. The comprehension of structural behaviour by studying the spectral features and the dynamics also supports the flexible nature of the r(CAG) motif.
- Centre for Biosciences and Biomedical Engineering, Indian Institute of Technology Indore, Indore, Madhya Pradesh, India.
Organizational Affiliation: