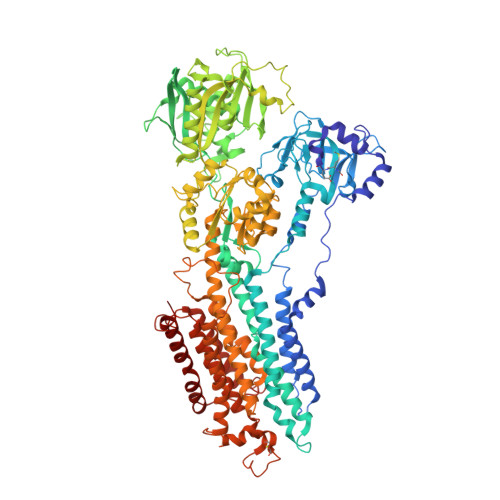
Biselyngbyasides, cytotoxic marine macrolides, are novel and potent inhibitors of the Ca(2+) pumps with a unique mode of binding
Morita, M., Ogawa, H., Ohno, O., Yamori, T., Suenaga, K., Toyoshima, C.(2015) FEBS Lett 589: 1406-1411
- PubMed: 25957767 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2015.04.056
- Primary Citation Related Structures:
4YCM, 4YCN - PubMed Abstract:
Biselyngbyasides (BLSs), macrolides from a marine cyanobacterium, are cytotoxic natural products whose target molecule is unknown. Here we report that BLSs are high affinity (Ki∼10 nM) inhibitors of Ca(2+)-pumps with a unique binding mode. The crystal structures of the Ca(2+)-pump in complex with BLSs at 3.2-3.5 Å-resolution show that BLSs bind to the pump near the cytoplasmic surface of the transmembrane region. The crystal structures and activity measurement of BLS analogs allow us to identify the structural features that confer high potency to BLSs as inhibitors of the pump.
- Department of Chemistry, Keio University, Kohoku-ku, Yokohama, Kanagawa 223-8522, Japan; Institute of Molecular and Cellular Biosciences, The University of Tokyo, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: