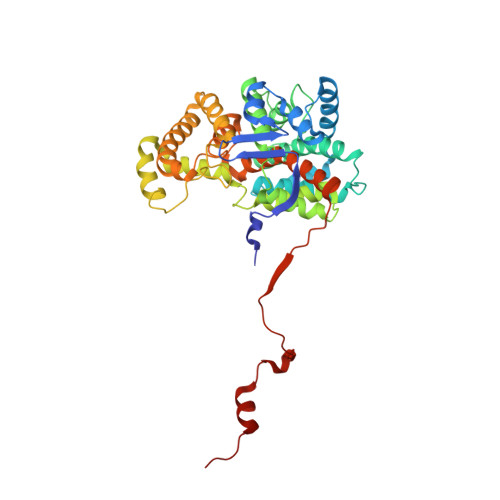
An active site-tail interaction in the structure of hexahistidine-tagged Thermoplasma acidophilum citrate synthase.
Murphy, J.R., Donini, S., Kappock, T.J.(2015) Acta Crystallogr F Struct Biol Commun 71: 1292-1299
- PubMed: 26457521 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15015939
- Primary Citation Related Structures:
4YBO - PubMed Abstract:
Citrate synthase (CS) plays a central metabolic role in aerobes and many other organisms. The CS reaction comprises two half-reactions: a Claisen aldol condensation of acetyl-CoA (AcCoA) and oxaloacetate (OAA) that forms citryl-CoA (CitCoA), and CitCoA hydrolysis. Protein conformational changes that `close' the active site play an important role in the assembly of a catalytically competent condensation active site. CS from the thermoacidophile Thermoplasma acidophilum (TpCS) possesses an endogenous Trp fluorophore that can be used to monitor the condensation reaction. The 2.2 Å resolution crystal structure of TpCS fused to a C-terminal hexahistidine tag (TpCSH6) reported here is an `open' structure that, when compared with several liganded TpCS structures, helps to define a complete path for active-site closure. One active site in each dimer binds a neighboring His tag, the first nonsubstrate ligand known to occupy both the AcCoA and OAA binding sites. Solution data collectively suggest that this fortuitous interaction is stabilized by the crystalline lattice. As a polar but almost neutral ligand, the active site-tail interaction provides a new starting point for the design of bisubstrate-analog inhibitors of CS.
- Department of Biochemistry, Purdue University, 175 South University Street, West Lafayette, IN 47907-2063, USA.
Organizational Affiliation: