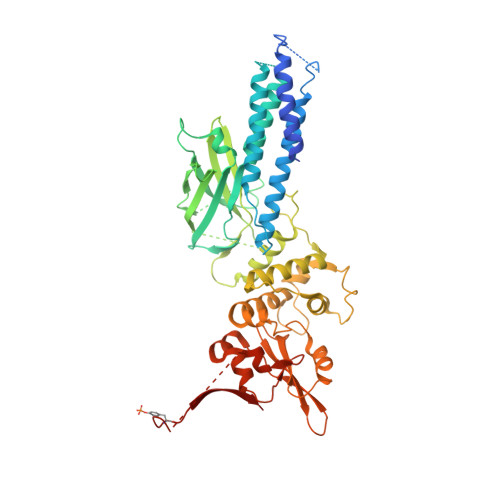
Structural basis for DNA recognition by STAT6
Li, J., Rodriguez, J.P., Niu, F., Pu, M., Wang, J., Hung, L.W., Shao, Q., Zhu, Y., Ding, W., Liu, Y., Da, Y., Yao, Z., Yang, J., Zhao, Y., Wei, G.H., Cheng, G., Liu, Z.J., Ouyang, S.(2016) Proc Natl Acad Sci U S A 113: 13015-13020
- PubMed: 27803324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1611228113
- Primary Citation Related Structures:
4Y5U, 4Y5W, 5D39 - PubMed Abstract:
STAT6 participates in classical IL-4/IL-13 signaling and stimulator of interferon genes-mediated antiviral innate immune responses. Aberrations in STAT6-mediated signaling are linked to development of asthma and diseases of the immune system. In addition, STAT6 remains constitutively active in multiple types of cancer. Therefore, targeting STAT6 is an attractive proposition for treating related diseases. Although a lot is known about the role of STAT6 in transcriptional regulation, molecular details on how STAT6 recognizes and binds specific segments of DNA to exert its function are not clearly understood. Here, we report the crystal structures of a homodimer of phosphorylated STAT6 core fragment (STAT6 CF ) alone and bound with the N3 and N4 DNA binding site. Analysis of the structures reveals that STAT6 undergoes a dramatic conformational change on DNA binding, which was further validated by performing molecular dynamics simulation studies and small angle X-ray scattering analysis. Our data show that a larger angle at the intersection where the two protomers of STAT meet and the presence of a unique residue, H415, in the DNA-binding domain play important roles in discrimination of the N4 site DNA from the N3 site by STAT6. H415N mutation of STAT6 CF decreased affinity of the protein for the N4 site DNA, but increased its affinity for N3 site DNA, both in vitro and in vivo. Results of our structure-function studies on STAT6 shed light on mechanism of DNA recognition by STATs in general and explain the reasons underlying STAT6's preference for N4 site DNA over N3.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: