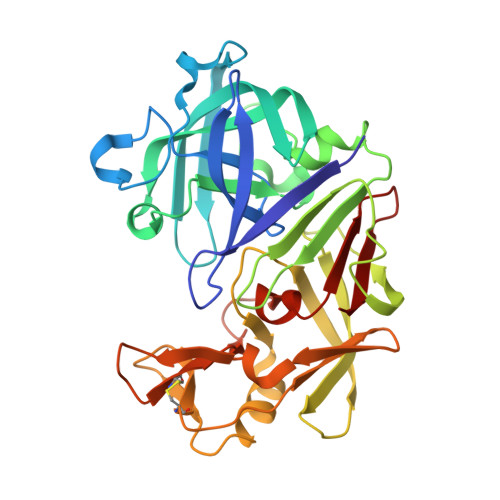
Structures of endothiapepsin-fragment complexes from crystallographic fragment screening using a novel, diverse and affordable 96-compound fragment library.
Huschmann, F.U., Linnik, J., Sparta, K., Uhlein, M., Wang, X., Metz, A., Schiebel, J., Heine, A., Klebe, G., Weiss, M.S., Mueller, U.(2016) Acta Crystallogr F Struct Biol Commun 72: 346-355
- PubMed: 27139825 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16004623
- Primary Citation Related Structures:
4Y38, 4Y3J, 4Y3Y, 4Y48, 4Y4D, 4Y4G, 4Y4J, 4ZE6, 4ZEA - PubMed Abstract:
Crystallographic screening of the binding of small organic compounds (termed fragments) to proteins is increasingly important for medicinal chemistry-oriented drug discovery. To enable such experiments in a widespread manner, an affordable 96-compound library has been assembled for fragment screening in both academia and industry. The library is selected from already existing protein-ligand structures and is characterized by a broad ligand diversity, including buffer ingredients, carbohydrates, nucleotides, amino acids, peptide-like fragments and various drug-like organic compounds. When applied to the model protease endothiapepsin in a crystallographic screening experiment, a hit rate of nearly 10% was obtained. In comparison to other fragment libraries and considering that no pre-screening was performed, this hit rate is remarkably high. This demonstrates the general suitability of the selected compounds for an initial fragment-screening campaign. The library composition, experimental considerations and time requirements for a complete crystallographic fragment-screening campaign are discussed as well as the nine fully refined obtained endothiapepsin-fragment structures. While most of the fragments bind close to the catalytic centre of endothiapepsin in poses that have been observed previously, two fragments address new sites on the protein surface. ITC measurements show that the fragments bind to endothiapepsin with millimolar affinity.
- Macromolecular Crystallography, Helmholtz-Zentrum Berlin, Albert-Einstein-Strasse 15, D-12489 Berlin, Germany.
Organizational Affiliation: