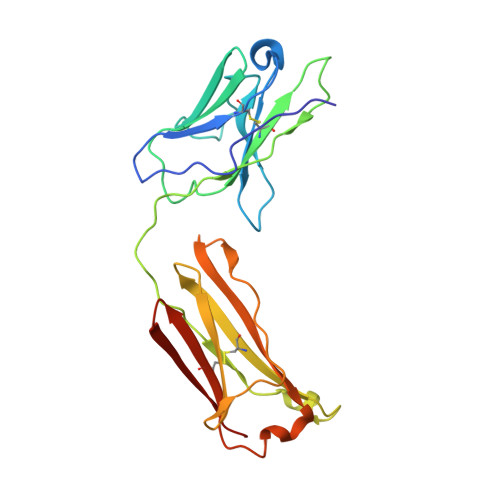
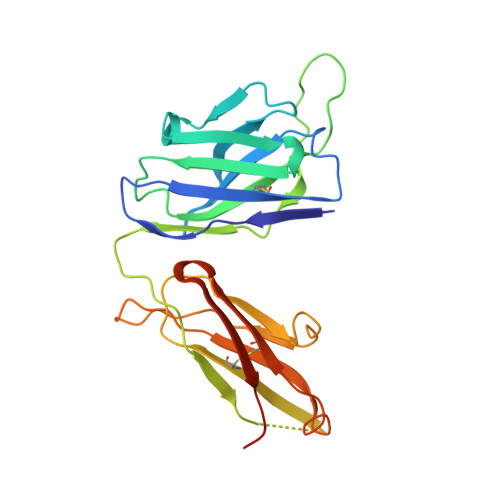
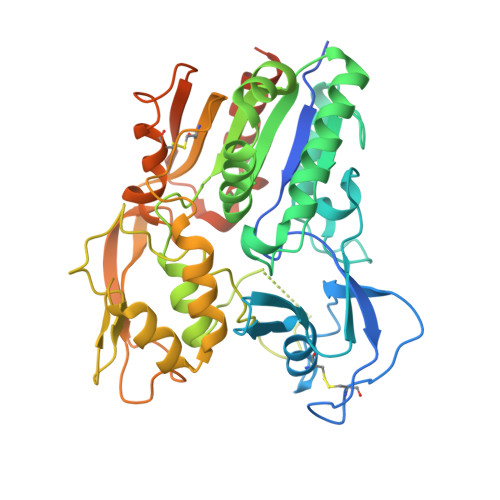
The high-resolution crystal structure of human LCAT.
Piper, D.E., Romanow, W.G., Gunawardane, R.N., Fordstrom, P., Masterman, S., Pan, O., Thibault, S.T., Zhang, R., Meininger, D., Schwarz, M., Wang, Z., King, C., Zhou, M., Walker, N.P.(2015) J Lipid Res 56: 1711-1719
- PubMed: 26195816 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1194/jlr.M059873
- Primary Citation Related Structures:
4XWG, 4XX1 - PubMed Abstract:
LCAT is intimately involved in HDL maturation and is a key component of the reverse cholesterol transport (RCT) pathway which removes excess cholesterol molecules from the peripheral tissues to the liver for excretion. Patients with loss-of-function LCAT mutations exhibit low levels of HDL cholesterol and corneal opacity. Here we report the 2.65 Å crystal structure of the human LCAT protein. Crystallization required enzymatic removal of N-linked glycans and complex formation with a Fab fragment from a tool antibody. The crystal structure reveals that LCAT has an α/β hydrolase core with two additional subdomains that play important roles in LCAT function. Subdomain 1 contains the region of LCAT shown to be required for interfacial activation, while subdomain 2 contains the lid and amino acids that shape the substrate binding pocket. Mapping the naturally occurring mutations onto the structure provides insight into how they may affect LCAT enzymatic activity.
- Therapeutic Discovery Amgen Inc., South San Francisco, CA 94080.
Organizational Affiliation: