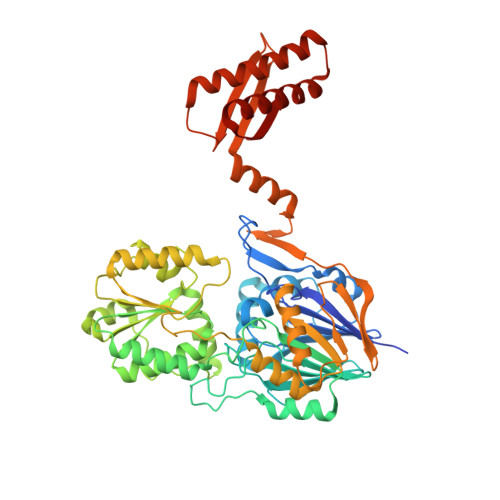
Structural insights into catalysis and dimerization enhanced exonuclease activity of RNase J
Zhao, Y., Lu, M., Zhang, H., Hu, J., Zhou, C., Xu, Q., Shah, A.M.U.H., Xu, H., Wang, L., Hua, Y.(2015) Nucleic Acids Res 43: 5550-5559
- PubMed: 25940620 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv444
- Primary Citation Related Structures:
4XWT, 4XWW - PubMed Abstract:
RNase J is a conserved ribonuclease that belongs to the β-CASP family of nucleases. It possesses both endo- and exo-ribonuclease activities, which play a key role in pre-rRNA maturation and mRNA decay. Here we report high-resolution crystal structures of Deinococcus radiodurans RNase J complexed with RNA or uridine 5'-monophosphate in the presence of manganese ions. Biochemical and structural studies revealed that RNase J uses zinc ions for two-metal-ion catalysis. One residue conserved among RNase J orthologues (motif B) forms specific electrostatic interactions with the scissile phosphate of the RNA that is critical for the catalysis and product stabilization. The additional manganese ion, which is coordinated by conserved residues at the dimer interface, is critical for RNase J dimerization and exonuclease activity. The structures may also shed light on the mechanism of RNase J exo- and endonucleolytic activity switch.
- Key Laboratory of Chinese Ministry of Agriculture for Nuclear-Agricultural Sciences, Institute of Nuclear-Agricultural Sciences, Zhejiang University, China yezhao@zju.edu.cn.
Organizational Affiliation: