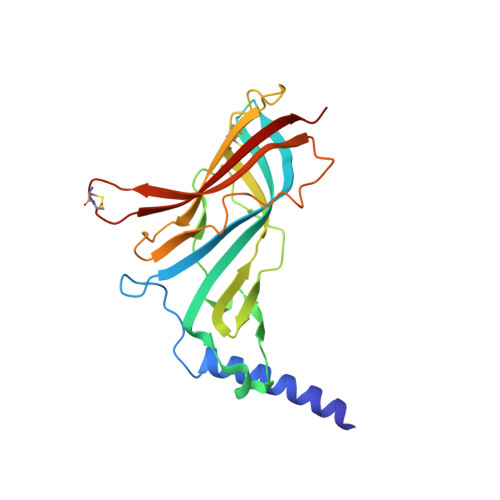
Marine Macrocyclic Imines, Pinnatoxins A and G: Structural Determinants and Functional Properties to Distinguish Neuronal alpha 7 from Muscle alpha 12 beta gamma delta nAChRs.
Bourne, Y., Sulzenbacher, G., Radic, Z., Araoz, R., Reynaud, M., Benoit, E., Zakarian, A., Servent, D., Molgo, J., Taylor, P., Marchot, P.(2015) Structure 23: 1106-1115
- PubMed: 26004441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.04.009
- Primary Citation Related Structures:
4XHE, 4XK9 - PubMed Abstract:
Pinnatoxins are macrocyclic imine phycotoxins associated with algal blooms and shellfish toxicity. Functional analysis of pinnatoxin A and pinnatoxin G by binding and voltage-clamp electrophysiology on membrane-embedded neuronal α7, α4β2, α3β2, and muscle-type α12βγδ nicotinic acetylcholine receptors (nAChRs) reveals high-affinity binding and potent antagonism for the α7 and α12βγδ subtypes. The toxins also bind to the nAChR surrogate, acetylcholine-binding protein (AChBP), with low Kd values reflecting slow dissociation. Crystal structures of pinnatoxin-AChBP complexes (1.9-2.2 Å resolution) show the multiple anchoring points of the hydrophobic portion, the cyclic imine, and the substituted bis-spiroketal and cyclohexene ring systems of the pinnatoxins that dictate tight binding between the opposing loops C and F at the receptor subunit interface, as observed for the 13-desmethyl-spirolide C and gymnodimine A congeners. Uniquely, however, the bulky bridged EF-ketal ring specific to the pinnatoxins extends radially from the interfacial-binding pocket to interact with the sequence-variable loop F and govern nAChR subtype selectivity and central neurotoxicity.
- Aix-Marseille Université, Laboratoire Architecture et Fonction des Macromolécules Biologiques, Campus Luminy, 13288 Marseille cedex 9, France; Centre National de la Recherche Scientifique, Laboratoire Architecture et Fonction des Macromolécules Biologiques, Campus Luminy, 13288 Marseille cedex 9, France. Electronic address: yves.bourne@afmb.univ-mrs.fr.
Organizational Affiliation: