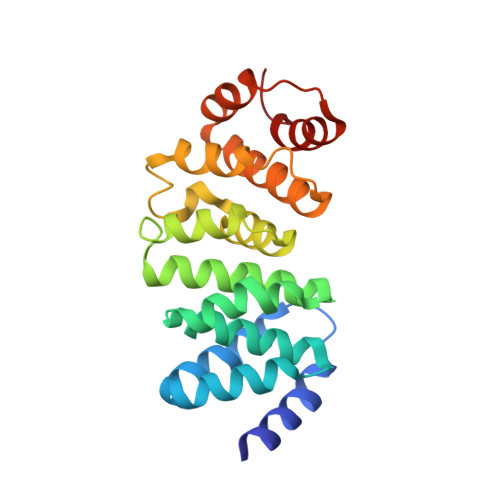
A New Family of HEAT-Like Repeat Proteins Lacking a Critical Substrate Recognition Motif Present in Related DNA Glycosylases.
Mullins, E.A., Shi, R., Kotsch, L.A., Eichman, B.F.(2015) PLoS One 10: e0127733-e0127733
- PubMed: 25978435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0127733
- Primary Citation Related Structures:
4X8Q - PubMed Abstract:
DNA glycosylases are important repair enzymes that eliminate a diverse array of aberrant nucleobases from the genomes of all organisms. Individual bacterial species often contain multiple paralogs of a particular glycosylase, yet the molecular and functional distinctions between these paralogs are not well understood. The recently discovered HEAT-like repeat (HLR) DNA glycosylases are distributed across all domains of life and are distinct in their specificity for cationic alkylpurines and mechanism of damage recognition. Here, we describe a number of phylogenetically diverse bacterial species with two orthologs of the HLR DNA glycosylase AlkD. One ortholog, which we designate AlkD2, is substantially less conserved. The crystal structure of Streptococcus mutans AlkD2 is remarkably similar to AlkD but lacks the only helix present in AlkD that penetrates the DNA minor groove. We show that AlkD2 possesses only weak DNA binding affinity and lacks alkylpurine excision activity. Mutational analysis of residues along this DNA binding helix in AlkD substantially reduced binding affinity for damaged DNA, for the first time revealing the importance of this structural motif for damage recognition by HLR glycosylases.
- Department of Biological Sciences and Center for Structural Biology, Vanderbilt University, Nashville, Tennessee, United States of America.
Organizational Affiliation: