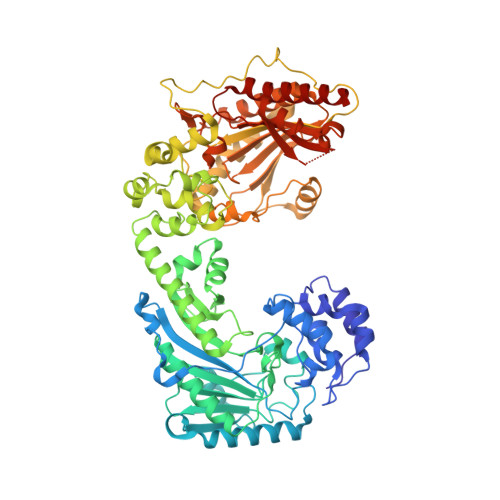
The Substrate-free and -bound Crystal Structures of the Duplicated Taurocyamine Kinase from the Human Parasite Schistosoma mansoni.
Merceron, R., Awama, A.M., Montserret, R., Marcillat, O., Gouet, P.(2015) J Biological Chem 290: 12951-12963
- PubMed: 25837252 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.628909
- Primary Citation Related Structures:
4WO8, 4WOD, 4WOE - PubMed Abstract:
The taurocyamine kinase from the blood fluke Schistosoma mansoni (SmTK) belongs to the phosphagen kinase (PK) family and catalyzes the reversible Mg(2+)-dependent transfer of a phosphoryl group between ATP and taurocyamine. SmTK is derived from gene duplication, as are all known trematode TKs. Our crystallographic study of SmTK reveals the first atomic structure of both a TK and a PK with a bilobal structure. The two unliganded lobes present a canonical open conformation and interact via their respective C- and N-terminal domains at a helix-mediated interface. This spatial arrangement differs from that observed in true dimeric PKs, in which both N-terminal domains make contact. Our structures of SmTK complexed with taurocyamine or l-arginine compounds explain the mechanism by which an arginine residue of the phosphagen specificity loop is crucial for substrate specificity. An SmTK crystal was soaked with the dead end transition state analog (TSA) components taurocyamine-NO3 (2-)-MgADP. One SmTK monomer was observed with two bound TSAs and an asymmetric conformation, with the first lobe semiclosed and the second closed. However, isothermal titration calorimetry and enzyme kinetics experiments showed that the two lobes function independently. A small angle x-ray scattering model of SmTK-TSA in solution with two closed active sites was generated.
- From the Institut de Biologie et Chimie des Protéines, BMSSI-IBCP, UMR 5086 CNRS Université Lyon 1, 7, Passage du Vercors, 69367 Lyon Cedex 07, France and.
Organizational Affiliation: