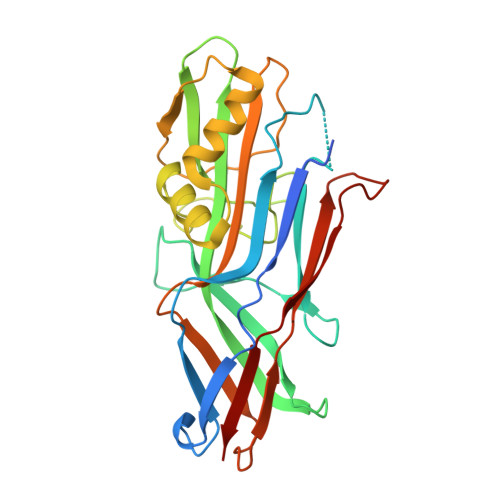
Structural and Functional Insight into the Carbohydrate Receptor Binding of F4 Fimbriae-producing Enterotoxigenic Escherichia coli.
Moonens, K., Van den Broeck, I., De Kerpel, M., Deboeck, F., Raymaekers, H., Remaut, H., De Greve, H.(2015) J Biological Chem 290: 8409-8419
- PubMed: 25631050 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.618595
- Primary Citation Related Structures:
4WE2, 4WEI - PubMed Abstract:
Enterotoxigenic Escherichia coli (ETEC) strains are important causes of intestinal disease in humans and lead to severe production losses in animal farming. A range of fimbrial adhesins in ETEC strains determines host and tissue tropism. ETEC strains expressing F4 fimbriae are associated with neonatal and post-weaning diarrhea in piglets. Three naturally occurring variants of F4 fimbriae (F4ab, F4ac, and F4ad) exist that differ in the primary sequence of their major adhesive subunit FaeG, and each features a related yet distinct receptor binding profile. Here the x-ray structure of FaeGad bound to lactose provides the first structural insight into the receptor specificity and mode of binding by the poly-adhesive F4 fimbriae. A small D'-D″-α1-α2 subdomain grafted on the immunoglobulin-like core of FaeG hosts the carbohydrate binding site. Two short amino acid stretches Phe(150)-Glu(152) and Val(166)-Glu(170) of FaeGad bind the terminal galactose in the lactosyl unit and provide affinity and specificity to the interaction. A hemagglutination-based assay with E. coli expressing mutant F4ad fimbriae confirmed the elucidated co-complex structure. Interestingly, the crucial D'-α1 loop that borders the FaeGad binding site adopts a different conformation in the two other FaeG variants and hints at a heterogeneous binding pocket among the FaeG serotypes.
- From the Structural and Molecular Microbiology, VIB Structural Biology Research Center, 1050 Brussels, the Structural Biology Brussels, Vrije Universiteit Brussel, Pleinlaan 2, 1050 Brussels, and.
Organizational Affiliation: