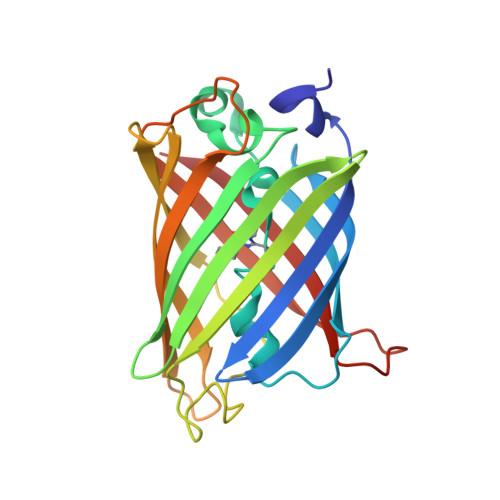
A Suite of Engineered GFP Molecules for Oligomeric Scaffolding.
Leibly, D.J., Arbing, M.A., Pashkov, I., DeVore, N., Waldo, G.S., Terwilliger, T.C., Yeates, T.O.(2015) Structure 23: 1754-1768
- PubMed: 26278175 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.07.008
- Primary Citation Related Structures:
4W69, 4W6A, 4W6B, 4W6C, 4W6D, 4W6F, 4W6G, 4W6H, 4W6I, 4W6J, 4W6K, 4W6L, 4W6M, 4W6N, 4W6O, 4W6P, 4W6R, 4W6S, 4W6T, 4W6U, 4W72, 4W73, 4W74, 4W75, 4W76, 4W77, 4W7A, 4W7C, 4W7D, 4W7E, 4W7F, 4W7R, 4W7X - PubMed Abstract:
Applications ranging from synthetic biology to protein crystallization could be advanced by facile systems for connecting multiple proteins together in predefined spatial relationships. One approach to this goal is to engineer many distinct assembly forms of a single carrier protein or scaffold, to which other proteins of interest can then be readily attached. In this work we chose GFP as a scaffold and engineered many alternative oligomeric forms, driven by either specific disulfide bond formation or metal ion addition. We generated a wide range of spatial arrangements of GFP subunits from 11 different oligomeric variants, and determined their X-ray structures in a total of 33 distinct crystal forms. Some of the oligomeric GFP variants show geometric polymorphism depending on conditions, while others show considerable geometric rigidity. Potential future applications of this system are discussed.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, CA 90095, USA; UCLA-DOE Institute of Genomics and Proteomics, University of California, Los Angeles, CA 90095, USA.
Organizational Affiliation: