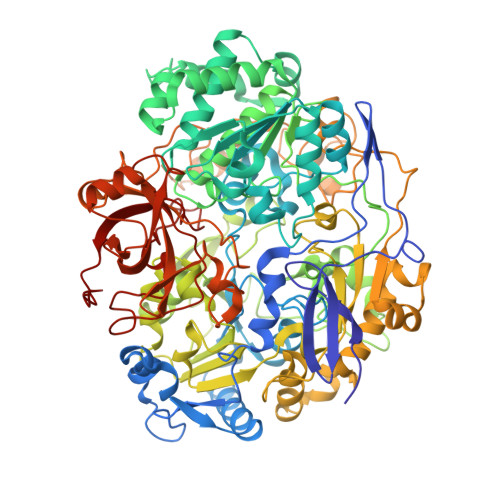
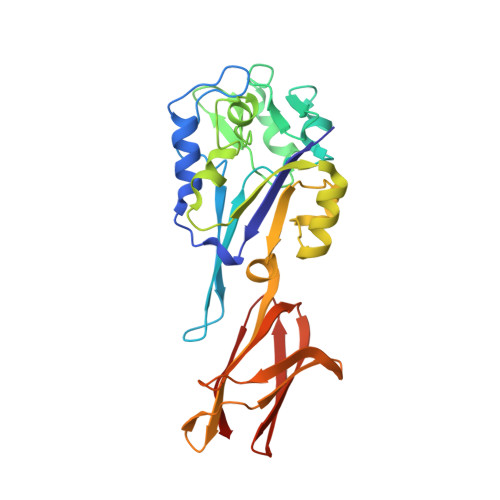
Crystal structure of pyrogallol-phloroglucinol transhydroxylase, an Mo enzyme capable of intermolecular hydroxyl transfer between phenols
Messerschmidt, A., Niessen, H., Abt, D., Einsle, O., Schink, B., Kroneck, P.M.H.(2004) Proc Natl Acad Sci U S A 101: 11571-11576
- PubMed: 15284442 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0404378101
- Primary Citation Related Structures:
4V4C, 4V4D, 4V4E - PubMed Abstract:
The Mo enzyme transhydroxylase from the anaerobic microorganism Pelobacter acidigallici catalyzes the conversion of pyrogallol to phloroglucinol. Such trihydroxybenzenes and their derivatives represent important building blocks of plant polymers. None of the transferred hydroxyl groups originates from water during transhydroxylation; instead a cosubstrate, such as 1,2,3,5-tetrahydroxybenzene, is used in a reaction without apparent electron transfer. Here, we report on the crystal structure of the enzyme in the reduced Mo(IV) state, which we solved by single anomalous-diffraction technique. It represents the largest structure (1,149 amino acid residues per molecule, 12 independent molecules per unit cell), which has been solved so far by single anomalous-diffraction technique. Tranhydroxylase is a heterodimer, with the active Mo-molybdopterin guanine dinucleotide (MGD)(2) site in the alpha-subunit, and three [4Fe-4S] centers in the beta-subunit. The latter subunit carries a seven-stranded, mainly antiparallel beta-barrel domain. We propose a scheme for the transhydroxylation reaction based on 3D structures of complexes of the enzyme with various polyphenols serving either as substrate or inhibitor.
- Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, Am Klopferspitz 18, 82152 Martinsried, Germany.
Organizational Affiliation: