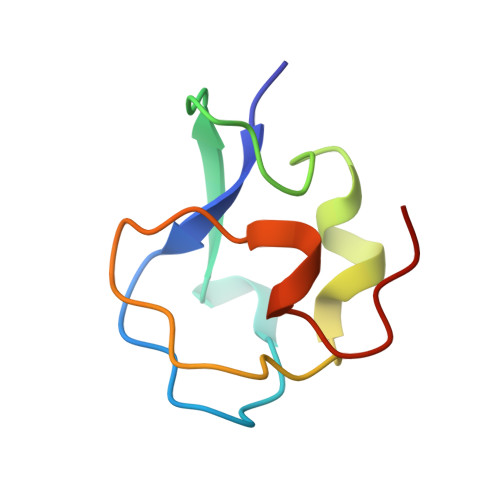
Purification, Crystal Structure Determination and Functional Characterization of Type III Antifreeze Proteins from the European Eelpout Zoarces Viviparus.
Wilkens, C., Poulsen, J.N., Ramlov, H., Lo Leggio, L.(2014) Cryobiology 69: 163
- PubMed: 25025819 Search on PubMed
- DOI: https://doi.org/10.1016/j.cryobiol.2014.07.003
- Primary Citation Related Structures:
4UR4, 4UR6 - PubMed Abstract:
Antifreeze proteins (AFPs) are essential components of many organisms adaptation to cold temperatures. Fish type III AFPs are divided into two groups, SP isoforms being much less active than QAE1 isoforms. Two type III AFPs from Zoarces viviparus, a QAE1 (ZvAFP13) and an SP (ZvAFP6) isoform, are here characterized and their crystal structures determined. We conclude that the higher activity of the QAE1 isoforms cannot be attributed to single residues, but rather a combination of structural effects. Furthermore both ZvAFP6 and ZvAFP13 crystal structures have water molecules around T18 equivalent to the tetrahedral-like waters previously identified in a neutron crystal structure. Interestingly, ZvAFP6 forms dimers in the crystal, with a significant dimer interface. The presence of ZvAFP6 dimers was confirmed in solution by native electrophoresis and gel filtration. To our knowledge this is the first report of dimerization of AFP type III proteins.
- Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark. Electronic address: cwil@bio.dtu.dk.
Organizational Affiliation: