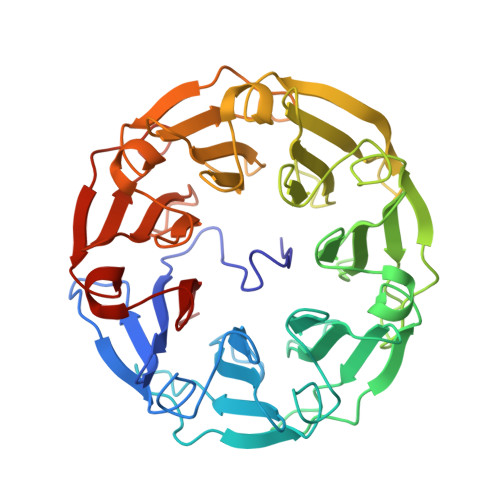
A Recombinant Fungal Lectin for Labeling Truncated Glycans on Human Cancer Cells.
Audfray, A., Beldjoudi, M., Breiman, A., Hurbin, A., Boos, I., Unverzagt, C., Bouras, M., Lantuejoul, S., Coll, J., Varrot, A., Le Pendu, J., Busser, B., Imberty, A.(2015) PLoS One 10: 01281
- PubMed: 26042789 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0128190
- Primary Citation Related Structures:
4UP4 - PubMed Abstract:
Cell surface glycoconjugates present alterations of their structures in chronic diseases and distinct oligosaccharide epitopes have been associated with cancer. Among them, truncated glycans present terminal non-reducing β-N-acetylglucosamine (GlcNAc) residues that are rare on healthy tissues. Lectins from unconventional sources such as fungi or algi provide novel markers that bind specifically to such epitopes, but their availability may be challenging. A GlcNAc-binding lectin from the fruiting body of the fungus Psathyrella velutina (PVL) has been produced in good yield in bacterial culture. A strong specificity for terminal GlcNAc residues was evidenced by glycan array. Affinity values obtained by microcalorimetry and surface plasmon resonance demonstrated a micromolar affinity for GlcNAcβ1-3Gal epitopes and for biantennary N-glycans with GlcNAcβ1-2Man capped branches. Crystal structure of PVL complexed with GlcNAcβ1-3Gal established the structural basis of the specificity. Labeling of several types of cancer cells and use of inhibitors of glycan metabolism indicated that rPVL binds to terminal GlcNAc but also to sialic acid (Neu5Ac). Analysis of glycosyltransferase expression confirmed the higher amount of GlcNAc present on cancer cells. rPVL binding is specific to cancer tissue and weak or no labeling is observed for healthy ones, except for stomach glands that present unique αGlcNAc-presenting mucins. In lung, breast and colon carcinomas, a clear delineation could be observed between cancer regions and surrounding healthy tissues. PVL is therefore a useful tool for labeling agalacto-glycans in cancer or other diseases.
- CERMAV, UPR5301, CNRS, University Grenoble Alpes, 38041 Grenoble, France.
Organizational Affiliation: