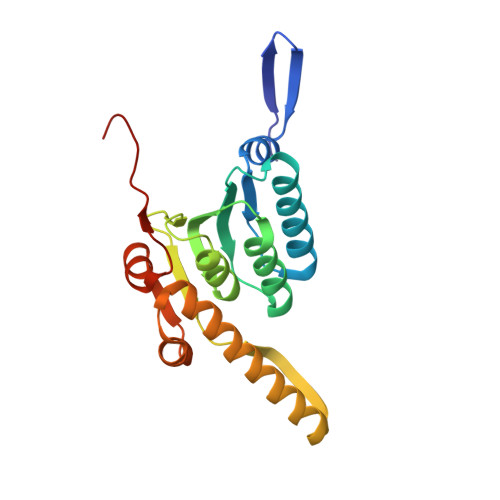
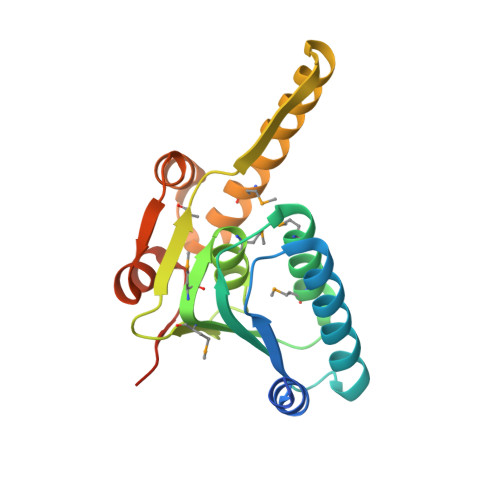
Crystal structure of Mycobacterium tuberculosis ClpP1P2 suggests a model for peptidase activation by AAA+ partner binding and substrate delivery.
Schmitz, K.R., Carney, D.W., Sello, J.K., Sauer, R.T.(2014) Proc Natl Acad Sci U S A 111: E4587-E4595
- PubMed: 25267638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1417120111
- Primary Citation Related Structures:
4U0G, 4U0H - PubMed Abstract:
Caseinolytic peptidase P (ClpP), a double-ring peptidase with 14 subunits, collaborates with ATPases associated with diverse activities (AAA+) partners to execute ATP-dependent protein degradation. Although many ClpP enzymes self-assemble into catalytically active homo-tetradecamers able to cleave small peptides, the Mycobacterium tuberculosis enzyme consists of discrete ClpP1 and ClpP2 heptamers that require a AAA+ partner and protein-substrate delivery or a peptide agonist to stabilize assembly of the active tetradecamer. Here, we show that cyclic acyldepsipeptides (ADEPs) and agonist peptides synergistically activate ClpP1P2 by mimicking AAA+ partners and substrates, respectively, and determine the structure of the activated complex. Our studies establish the basis of heteromeric ClpP1P2 assembly and function, reveal tight coupling between the conformations of each ring, show that ADEPs bind only to one ring but appear to open the axial pores of both rings, provide a foundation for rational drug development, and suggest strategies for studying the roles of individual ClpP1 and ClpP2 rings in Clp-family proteolysis.
- Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139; and.
Organizational Affiliation: