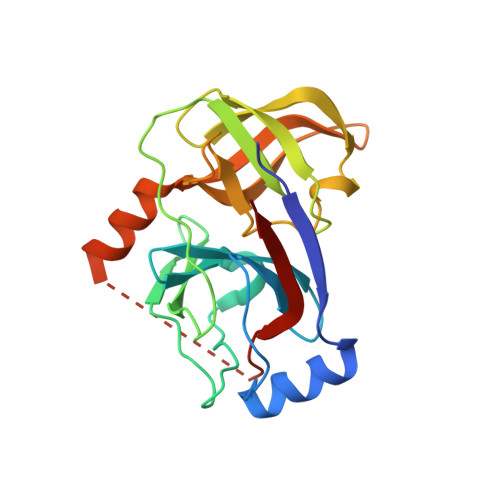
Structure-based design of a novel series of azetidine inhibitors of the hepatitis C virus NS3/4A serine protease.
Parsy, C., Alexandre, F.R., Brandt, G., Caillet, C., Cappelle, S., Chaves, D., Convard, T., Derock, M., Gloux, D., Griffon, Y., Lallos, L., Leroy, F., Liuzzi, M., Loi, A.G., Moulat, L., Musiu, C., Rahali, H., Roques, V., Seifer, M., Standring, D., Surleraux, D.(2014) Bioorg Med Chem Lett 24: 4444-4449
- PubMed: 25155387 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2014.08.002
- Primary Citation Related Structures:
4TYD - PubMed Abstract:
Structural homology between thrombin inhibitors and the early tetrapeptide HCV protease inhibitor led to the bioisosteric replacement of the P2 proline by a 2,4-disubstituted azetidine within the macrocyclic β-strand mimic. Molecular modeling guided the design of the series. This approach was validated by the excellent activity and selectivity in biochemical and cell based assays of this novel series and confirmed by the co-crystal structure of the inhibitor with the NS3/4A protein (PDB code: 4TYD).
- IDENIX Pharmaceuticals, 1682 rue de la Valsière, Cap Gamma, BP 50001, 34189 Montpellier Cedex 4, France. Electronic address: parsy.christophe@idenix.com.
Organizational Affiliation: