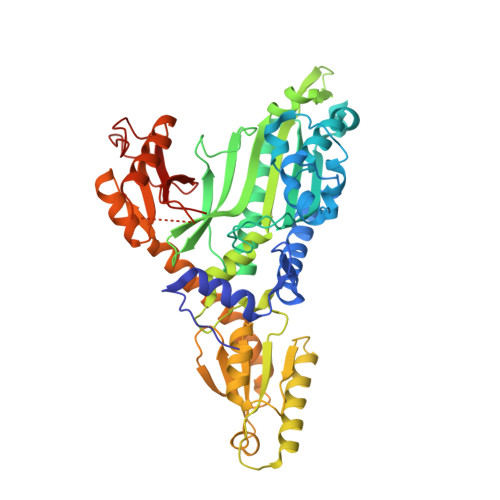
Structural and functional analysis of the anti-malarial drug target prolyl-tRNA synthetase.
Jain, V., Kikuchi, H., Oshima, Y., Sharma, A., Yogavel, M.(2014) J Struct Funct Genomics 15: 181-190
- PubMed: 25047712 Search on PubMed
- DOI: https://doi.org/10.1007/s10969-014-9186-x
- Primary Citation Related Structures:
4TWA - PubMed Abstract:
Aminoacyl-tRNA synthetases (aaRSs) drive protein translation in cells and hence these are essential enzymes across life. Inhibition of these enzymes can halt growth of an organism by stalling protein translation. Therefore, small molecule targeting of aaRS active sites is an attractive avenue from the perspective of developing anti-infectives. Febrifugine and its derivatives like halofuginone (HF) are known to inhibit prolyl-tRNA synthetase of malaria parasite Plasmodium falciparum. Here, we present functional and crystallographic data on P. falciparum prolyl-tRNA synthetase (PfPRS). Using immunofluorescence data, we show that PfPRS is exclusively resident in the parasite cytoplasm within asexual blood stage parasites. The inhibitor HF interacts strongly with PfPRS in a non-competitive binding mode in presence or absence of ATP analog. Intriguingly, the two monomers that constitute dimeric PfPRS display significantly different conformations in their active site regions. The structural analyses presented here provide a framework for development of febrifugine derivatives that can seed development of new anti-malarials.
- Structural and Computational Biology Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), New Delhi, 110067, India.
Organizational Affiliation: