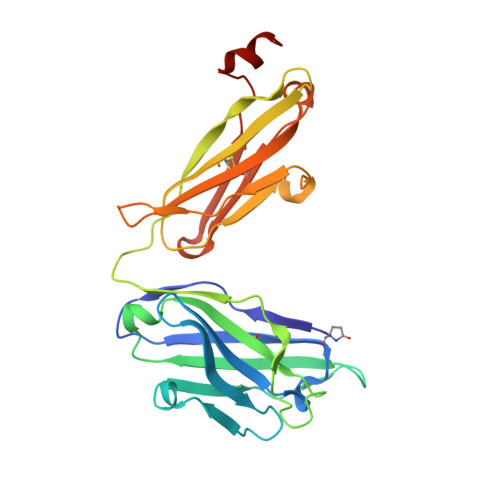
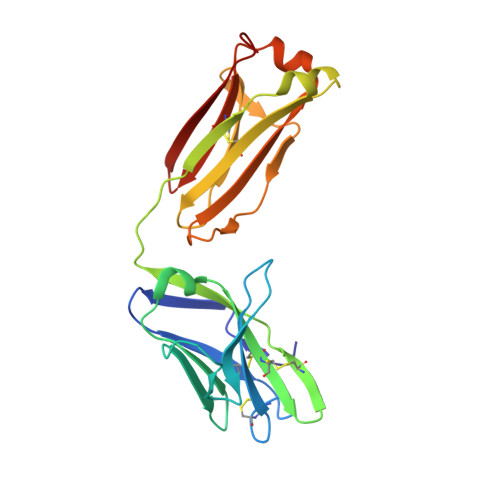
Broad and potent HIV-1 neutralization by a human antibody that binds the gp41-gp120 interface.
Huang, J., Kang, B.H., Pancera, M., Lee, J.H., Tong, T., Feng, Y., Imamichi, H., Georgiev, I.S., Chuang, G.Y., Druz, A., Doria-Rose, N.A., Laub, L., Sliepen, K., van Gils, M.J., de la Pena, A.T., Derking, R., Klasse, P.J., Migueles, S.A., Bailer, R.T., Alam, M., Pugach, P., Haynes, B.F., Wyatt, R.T., Sanders, R.W., Binley, J.M., Ward, A.B., Mascola, J.R., Kwong, P.D., Connors, M.(2014) Nature 515: 138-142
- PubMed: 25186731 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13601
- Primary Citation Related Structures:
4TOY - PubMed Abstract:
The isolation of human monoclonal antibodies is providing important insights into the specificities that underlie broad neutralization of HIV-1 (reviewed in ref. 1). Here we report a broad and extremely potent HIV-specific monoclonal antibody, termed 35O22, which binds a novel HIV-1 envelope glycoprotein (Env) epitope. 35O22 neutralized 62% of 181 pseudoviruses with a half-maximum inhibitory concentration (IC50) <50 μg ml(-1). The median IC50 of neutralized viruses was 0.033 μg ml(-1), among the most potent thus far described. 35O22 did not bind monomeric forms of Env tested, but did bind the trimeric BG505 SOSIP.664. Mutagenesis and a reconstruction by negative-stain electron microscopy of the Fab in complex with trimer revealed that it bound to a conserved epitope, which stretched across gp120 and gp41. The specificity of 35O22 represents a novel site of vulnerability on HIV Env, which serum analysis indicates to be commonly elicited by natural infection. Binding to this new site of vulnerability may thus be an important complement to current monoclonal-antibody-based approaches to immunotherapies, prophylaxis and vaccine design.
- Laboratory of Immunoregulation, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: