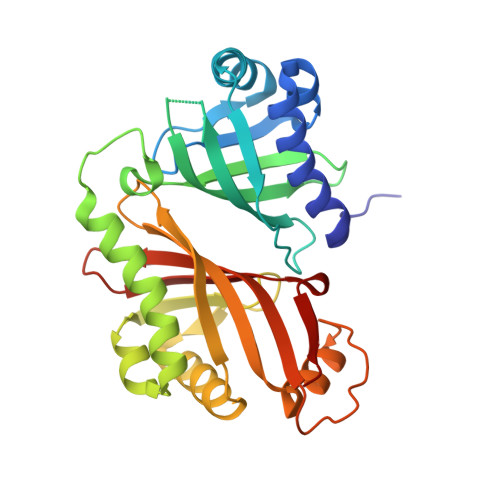
Epoxide hydrolase-lasalocid a structure provides mechanistic insight into polyether natural product biosynthesis.
Wong, F.T., Hotta, K., Chen, X., Fang, M., Watanabe, K., Kim, C.Y.(2015) J Am Chem Soc 137: 86-89
- PubMed: 25535803 Search on PubMed
- DOI: https://doi.org/10.1021/ja511374k
- Primary Citation Related Structures:
4RZM - PubMed Abstract:
Biosynthesis of some polyether natural products involves a kinetically disfavored epoxide-opening cyclic ether formation, a reaction termed anti-Baldwin cyclization. One such example is the biosynthesis of lasalocid A, an ionophore antibiotic polyether. During lasalocid A biosynthesis, an epoxide hydrolase, Lsd19, converts the bisepoxy polyketide intermediate into the tetrahydrofuranyl-tetrahydropyran product. We report the crystal structure of Lsd19 in complex with lasalocid A. The structure unambiguously shows that the C-terminal domain of Lsd19 catalyzes the intriguing anti-Baldwin cyclization. We propose a general mechanism for epoxide selection by ionophore polyether epoxide hydrolases.
- Department of Biological Sciences, National University of Singapore , 117543 Singapore.
Organizational Affiliation: