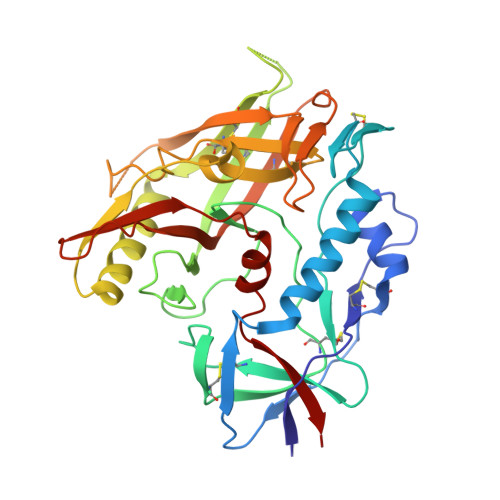
Structure-Based Design of a Small Molecule CD4-Antagonist with Broad Spectrum Anti-HIV-1 Activity.
Curreli, F., Kwon, Y.D., Zhang, H., Scacalossi, D., Belov, D.S., Tikhonov, A.A., Andreev, I.A., Altieri, A., Kurkin, A.V., Kwong, P.D., Debnath, A.K.(2015) J Med Chem 58: 6909-6927
- PubMed: 26301736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00709
- Primary Citation Related Structures:
4RZ8 - PubMed Abstract:
Earlier we reported the discovery and design of NBD-556 and their analogs which demonstrated their potential as HIV-1 entry inhibitors. However, progress in developing these inhibitors has been stymied by their CD4-agonist properties, an unfavorable trait for use as drug. Here, we demonstrate the successful conversion of a full CD4-agonist (NBD-556) through a partial CD4-agonist (NBD-09027), to a full CD4-antagonist (NBD-11021) by structure-based modification of the critical oxalamide midregion, previously thought to be intolerant of modification. NBD-11021 showed unprecedented neutralization breath for this class of inhibitors, with pan-neutralization against a panel of 56 Env-pseudotyped HIV-1 representing diverse subtypes of clinical isolates (IC50 as low as 270 nM). The cocrystal structure of NBD-11021 complexed to a monomeric HIV-1 gp120 core revealed its detail binding characteristics. The study is expected to provide a framework for further development of NBD series as HIV-1 entry inhibitors for clinical application against AIDS.
- Laboratory of Molecular Modeling and Drug Design, Lindsey F. Kimball Research Institute, New York Blood Center, New York, New York 10065, United States.
Organizational Affiliation: