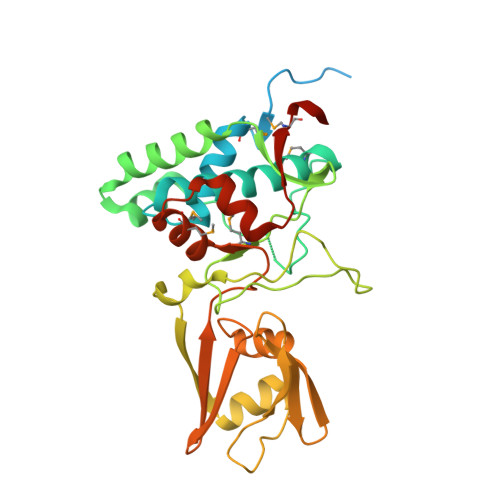
Crystal Structures of the SpoIID Lytic Transglycosylases Essential for Bacterial Sporulation.
Nocadello, S., Minasov, G., Shuvalova, L.S., Dubrovska, I., Sabini, E., Anderson, W.F.(2016) J Biological Chem 291: 14915-14926
- PubMed: 27226615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.729749
- Primary Citation Related Structures:
4RWR, 5I1T - PubMed Abstract:
Bacterial spores are the most resistant form of life known on Earth and represent a serious problem for (i) bioterrorism attack, (ii) horizontal transmission of microbial pathogens in the community, and (iii) persistence in patients and in a nosocomial environment. Stage II sporulation protein D (SpoIID) is a lytic transglycosylase (LT) essential for sporulation. The LT superfamily is a potential drug target because it is active in essential bacterial processes involving the peptidoglycan, which is unique to bacteria. However, the absence of structural information for the sporulation-specific LT enzymes has hindered mechanistic understanding of SpoIID. Here, we report the first crystal structures with and without ligands of the SpoIID family from two community relevant spore-forming pathogens, Bacillus anthracis and Clostridium difficile. The structures allow us to visualize the overall architecture, characterize the substrate recognition model, identify critical residues, and provide the structural basis for catalysis by this new family of enzymes.
- From the Center for Structural Genomics of Infectious Diseases, Department of Biochemistry and Molecular Genetics, Northwestern University Feinberg School of Medicine, Chicago, Illinois 60611.
Organizational Affiliation: