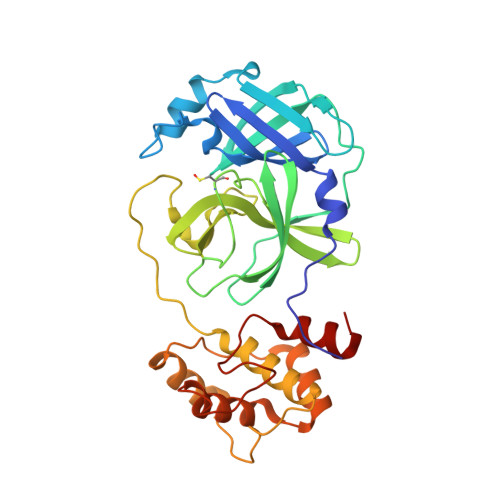
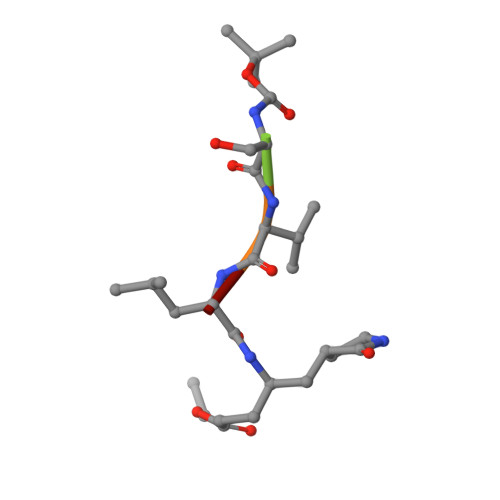
Ligand-induced Dimerization of Middle East Respiratory Syndrome (MERS) Coronavirus nsp5 Protease (3CLpro): IMPLICATIONS FOR nsp5 REGULATION AND THE DEVELOPMENT OF ANTIVIRALS.
Tomar, S., Johnston, M.L., St John, S.E., Osswald, H.L., Nyalapatla, P.R., Paul, L.N., Ghosh, A.K., Denison, M.R., Mesecar, A.D.(2015) J Biological Chem 290: 19403-19422
- PubMed: 26055715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.651463
- Primary Citation Related Structures:
4RSP, 4YLU - PubMed Abstract:
All coronaviruses, including the recently emerged Middle East respiratory syndrome coronavirus (MERS-CoV) from the β-CoV subgroup, require the proteolytic activity of the nsp5 protease (also known as 3C-like protease, 3CL(pro)) during virus replication, making it a high value target for the development of anti-coronavirus therapeutics. Kinetic studies indicate that in contrast to 3CL(pro) from other β-CoV 2c members, including HKU4 and HKU5, MERS-CoV 3CL(pro) is less efficient at processing a peptide substrate due to MERS-CoV 3CL(pro) being a weakly associated dimer. Conversely, HKU4, HKU5, and SARS-CoV 3CL(pro) enzymes are tightly associated dimers. Analytical ultracentrifugation studies support that MERS-CoV 3CL(pro) is a weakly associated dimer (Kd ∼52 μm) with a slow off-rate. Peptidomimetic inhibitors of MERS-CoV 3CL(pro) were synthesized and utilized in analytical ultracentrifugation experiments and demonstrate that MERS-CoV 3CL(pro) undergoes significant ligand-induced dimerization. Kinetic studies also revealed that designed reversible inhibitors act as activators at a low compound concentration as a result of induced dimerization. Primary sequence comparisons and x-ray structural analyses of two MERS-CoV 3CLpro and inhibitor complexes, determined to 1.6 Å, reveal remarkable structural similarity of the dimer interface with 3CL(pro) from HKU4-CoV and HKU5-CoV. Despite this structural similarity, substantial differences in the dimerization ability suggest that long range interactions by the nonconserved amino acids distant from the dimer interface may control MERS-CoV 3CL(pro) dimerization. Activation of MERS-CoV 3CL(pro) through ligand-induced dimerization appears to be unique within the genogroup 2c and may potentially increase the complexity in the development of MERS-CoV 3CL(pro) inhibitors as antiviral agents.
- From the Departments of Biological Sciences and.
Organizational Affiliation: