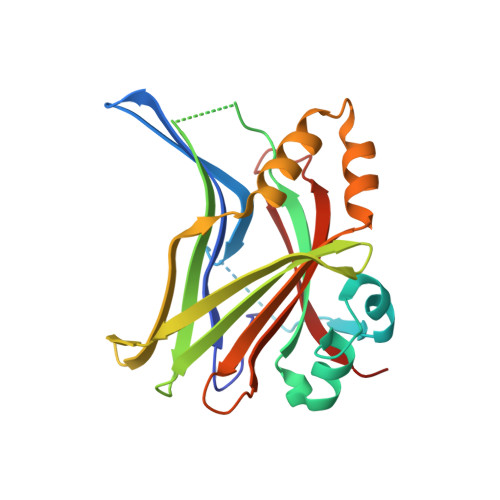
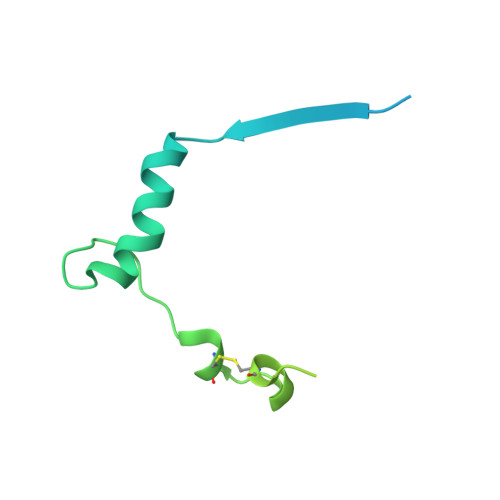
Targeting Hippo pathway by specific interruption of YAP-TEAD interaction using cyclic YAP-like peptides.
Zhou, Z., Hu, T., Xu, Z., Lin, Z., Zhang, Z., Feng, T., Zhu, L., Rong, Y., Shen, H., Luk, J.M., Zhang, X., Qin, N.(2015) FASEB J 29: 724-732
- PubMed: 25384421 Search on PubMed
- DOI: https://doi.org/10.1096/fj.14-262980
- Primary Citation Related Structures:
4RE1 - PubMed Abstract:
Hippo signaling pathway is emerging as a novel target for anticancer therapy because it plays key roles in organ size control and tumorigenesis. As the downstream effectors, Yes-associated protein (YAP)-transcriptional enhancer activation domain family member (TEAD) association is essential for YAP-driven oncogenic activity, while TEAD is largely dispensable for normal tissue growth. We present the design of YAP-like peptides (17mer) to occupy the interface 3 on TEAD. Introducing cysteines at YAP sites 87 and 96 can induce disulfide formation, as confirmed by crystallography. The engineered peptide significantly improves the potency in disrupting YAP-TEAD interaction in vitro. To confirm that blocking YAP-TEAD complex formation by directly targeting on TEAD is a valid approach, we report a significant reduction in tumor growth rate in a hepatocellular carcinoma xenograft model after introducing the dominant-negative mutation (Y406H) of TEAD1 to abolish YAP-TEAD interaction. Our results suggest that targeting TEAD is a promising strategy against YAP-induced oncogenesis.
- *Discovery Technology, Medicinal Chemistry, and Discovery Oncology, Roche Pharmaceutical Research and Early Development, Roche Innovation Center Shanghai, Shanghai, China joe.zhou.jz2@roche.com.
Organizational Affiliation: