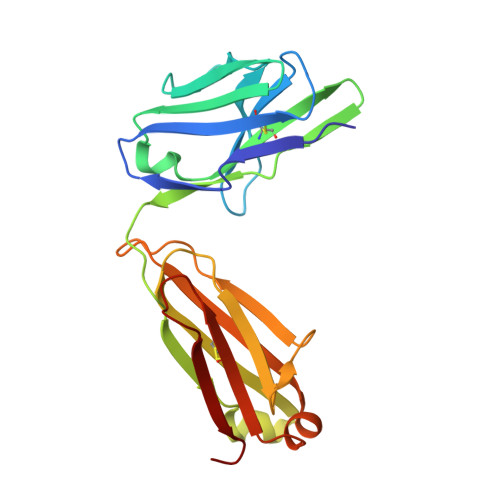
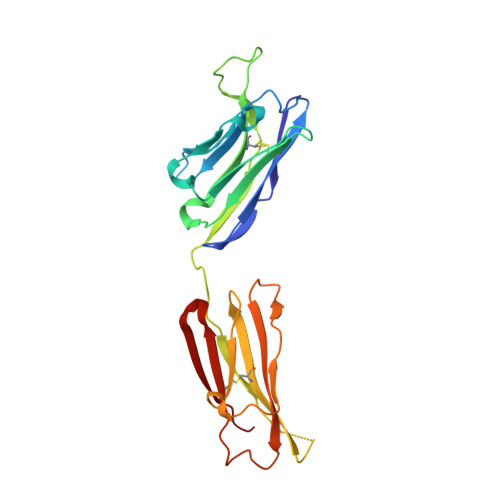
Crystal structure of the HIV neutralizing antibody 2G12 in complex with a bacterial oligosaccharide analog of mammalian oligomannose.
Stanfield, R.L., De Castro, C., Marzaioli, A.M., Wilson, I.A., Pantophlet, R.(2015) Glycobiology 25: 412-419
- PubMed: 25380763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/glycob/cwu123
- Primary Citation Related Structures:
4RBP - PubMed Abstract:
Human immunodeficiency virus-1 (HIV-1) is a major public health threat that continues to infect millions of people worldwide each year. A prophylactic vaccine remains the most cost-effective way of globally reducing and eliminating the spread of the virus. The HIV envelope spike, which is the target of many vaccine design efforts, is densely mantled with carbohydrate and several potent broadly neutralizing antibodies to HIV-1 recognize carbohydrate on the envelope spike as a major part of their epitope. However, immunizing with recombinant forms of the envelope glycoprotein does not typically elicit anti-carbohydrate antibodies. Thus, studies of alternative antigens that may serve as a starting point for carbohydrate-based immunogens are of interest. Here, we present the crystal structure of one such anti-carbohydrate HIV neutralizing antibody (2G12) in complex with the carbohydrate backbone of the lipooligosaccharide from Rhizobium radiobacter strain Rv3, which exhibits a chemical structure that naturally mimics the core high-mannose carbohydrate epitope of 2G12 on HIV-1 gp120. The structure described here provides molecular evidence of the structural homology between the Rv3 oligosaccharide and highly abundant carbohydrates on the surface of HIV-1 and raises the potential for the design of novel glycoconjugates that may find utility in efforts to develop immunogens for eliciting carbohydrate-specific neutralizing antibodies to HIV.
- Department of Integrative Structural and Computational Biology, Scripps CHAVI-ID, and IAVI Neutralizing Antibody Center, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: