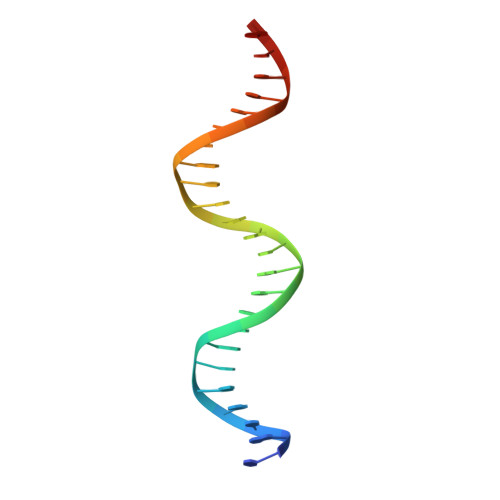
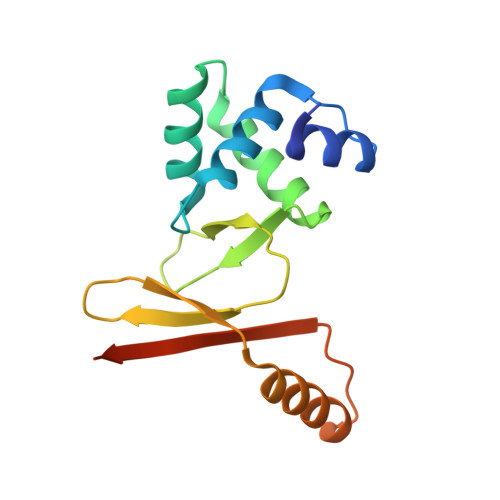
Mechanistic insights into metal ion activation and operator recognition by the ferric uptake regulator.
Deng, Z., Wang, Q., Liu, Z., Zhang, M., Machado, A.C., Chiu, T.P., Feng, C., Zhang, Q., Yu, L., Qi, L., Zheng, J., Wang, X., Huo, X., Qi, X., Li, X., Wu, W., Rohs, R., Li, Y., Chen, Z.(null) Nat Commun 6: 7642-7642
- PubMed: 26134419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8642
- Primary Citation Related Structures:
4RAY, 4RAZ, 4RB0, 4RB1, 4RB2, 4RB3 - PubMed Abstract:
Ferric uptake regulator (Fur) plays a key role in the iron homeostasis of prokaryotes, such as bacterial pathogens, but the molecular mechanisms and structural basis of Fur-DNA binding remain incompletely understood. Here, we report high-resolution structures of Magnetospirillum gryphiswaldense MSR-1 Fur in four different states: apo-Fur, holo-Fur, the Fur-feoAB1 operator complex and the Fur-Pseudomonas aeruginosa Fur box complex. Apo-Fur is a transition metal ion-independent dimer whose binding induces profound conformational changes and confers DNA-binding ability. Structural characterization, mutagenesis, biochemistry and in vivo data reveal that Fur recognizes DNA by using a combination of base readout through direct contacts in the major groove and shape readout through recognition of the minor-groove electrostatic potential by lysine. The resulting conformational plasticity enables Fur binding to diverse substrates. Our results provide insights into metal ion activation and substrate recognition by Fur that suggest pathways to engineer magnetotactic bacteria and antipathogenic drugs.
- State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing 100193, China.
Organizational Affiliation: