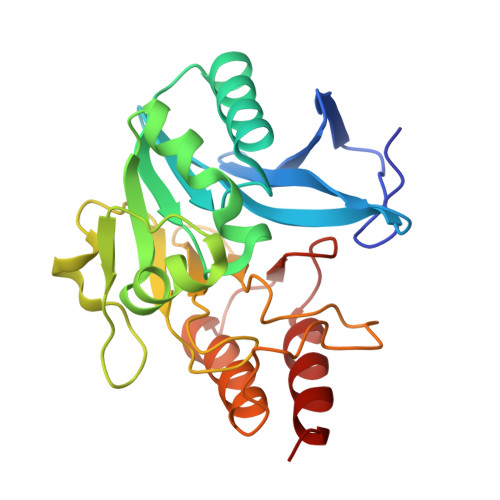
Crystal Structure of New Delhi Metallo-beta-Lactamase-1 Mutant M67V Complexed with Hydrolyzed Ampicillin
Kim, Y., Tesar, C., Jedrzejczak, R., Babnigg, G., Sacchettini, J., Joachimiak, A., Midwest Center for Structural Genomics (MCSG), Structures of Mtb Proteins Conferring Susceptibility to Known Mtb Inhibitors (MTBI)To be published.