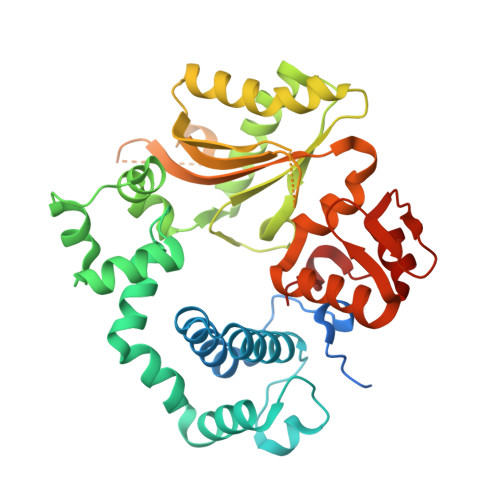
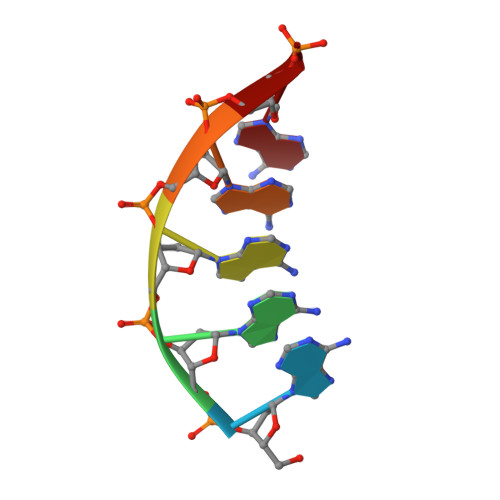
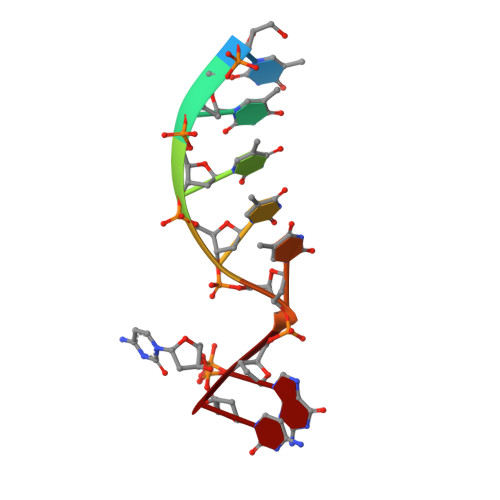
Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair.
Gouge, J., Rosario, S., Romain, F., Poitevin, F., Beguin, P., Delarue, M.(2015) EMBO J 34: 1126-1142
- PubMed: 25762590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.201489643
- Primary Citation Related Structures:
4QZ8, 4QZ9, 4QZA, 4QZB, 4QZC, 4QZD, 4QZE, 4QZF, 4QZG, 4QZH, 4QZI - PubMed Abstract:
Eukaryotic DNA polymerase mu of the PolX family can promote the association of the two 3'-protruding ends of a DNA double-strand break (DSB) being repaired (DNA synapsis) even in the absence of the core non-homologous end-joining (NHEJ) machinery. Here, we show that terminal deoxynucleotidyltransferase (TdT), a closely related PolX involved in V(D)J recombination, has the same property. We solved its crystal structure with an annealed DNA synapsis containing one micro-homology (MH) base pair and one nascent base pair. This structure reveals how the N-terminal domain and Loop 1 of Tdt cooperate for bridging the two DNA ends, providing a templating base in trans and limiting the MH search region to only two base pairs. A network of ordered water molecules is proposed to assist the incorporation of any nucleotide independently of the in trans templating base. These data are consistent with a recent model that explains the statistics of sequences synthesized in vivo by Tdt based solely on this dinucleotide step. Site-directed mutagenesis and functional tests suggest that this structural model is also valid for Pol mu during NHEJ.
- Unité de Dynamique Structurale des Macromolécules, Institut Pasteur, UMR 3528 du C.N.R.S., Paris, France.
Organizational Affiliation: