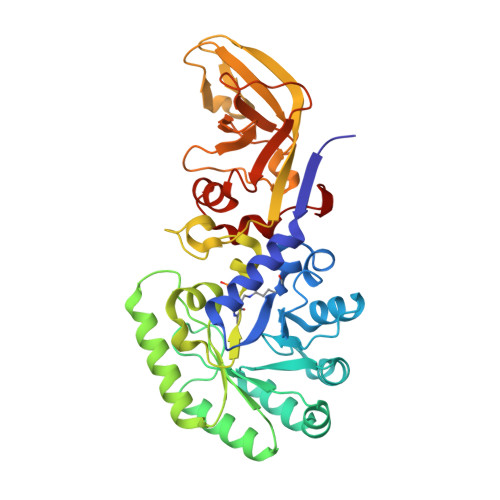
The structure of alanine racemase from Acinetobacter baumannii
Davis, E., Scaletti-Hutchinson, E., Opel-Reading, H., Nakatani, Y., Krause, K.L.(2014) Acta Crystallogr F Struct Biol Commun 70: 1199-1205
- PubMed: 25195891 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14017725
- Primary Citation Related Structures:
4QHR - PubMed Abstract:
Acinetobacter baumannii is an opportunistic Gram-negative bacterium which is a common cause of hospital-acquired infections. Numerous antibiotic-resistant strains exist, emphasizing the need for the development of new antimicrobials. Alanine racemase (Alr) is a pyridoxal 5'-phosphate dependent enzyme that is responsible for racemization between enantiomers of alanine. As D-alanine is an essential component of the bacterial cell wall, its inhibition is lethal to prokaryotes, making it an excellent antibiotic drug target. The crystal structure of A. baumannii alanine racemase (AlrAba) from the highly antibiotic-resistant NCTC13302 strain has been solved to 1.9 Å resolution. Comparison of AlrAba with alanine racemases from closely related bacteria demonstrates a conserved overall fold. The substrate entryway and active site of the enzymes were shown to be highly conserved. The structure of AlrAba will provide the template required for future structure-based drug-design studies.
- Department of Biochemistry, University of Otago, Dunedin, New Zealand.
Organizational Affiliation: