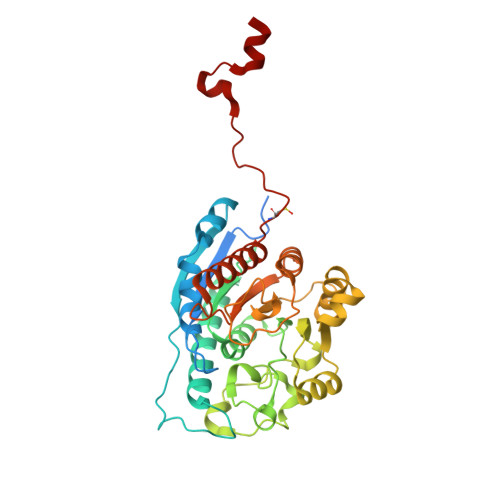
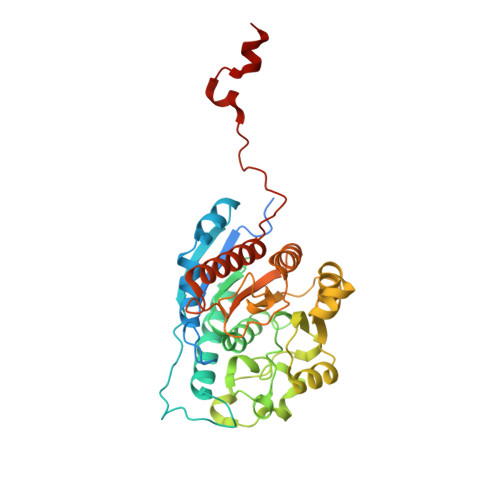
Crystal Structure of Schistosoma mansoni Arginase, a Potential Drug Target for the Treatment of Schistosomiasis.
Hai, Y., Edwards, J.E., Van Zandt, M.C., Hoffmann, K.F., Christianson, D.W.(2014) Biochemistry 53: 4671-4684
- PubMed: 25007099 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi5004519
- Primary Citation Related Structures:
4Q3P, 4Q3Q, 4Q3R, 4Q3S, 4Q3T, 4Q3U, 4Q3V, 4Q40, 4Q41, 4Q42 - PubMed Abstract:
The X-ray crystal structure of arginase from Schistosoma mansoni (SmARG) and the structures of its complexes with several amino acid inhibitors have been determined at atomic resolution. SmARG is a binuclear manganese metalloenzyme that catalyzes the hydrolysis of l-arginine to form l-ornithine and urea, and this enzyme is upregulated in all forms of the parasite that interact with the human host. Current hypotheses suggest that parasitic arginases could play a role in host immune evasion by depleting pools of substrate l-arginine that would otherwise be utilized for NO biosynthesis and NO-dependent processes in the immune response. Although the amino acid sequence of SmARG is only 42% identical with that of human arginase I, residues important for substrate binding and catalysis are strictly conserved. In general, classical amino acid inhibitors such as 2(S)-amino-6-boronohexanoic acid (ABH) tend to bind more weakly to SmARG than to human arginase I despite identical inhibitor binding modes in each enzyme active site. The identification of a patch on the enzyme surface capable of accommodating the additional Cα substitutent of an α,α-disubstituted amino acid inhibitor suggests that such inhibitors could exhibit higher affinity and biological activity. The structures of SmARG complexed with two different α,α-disubstituted derivatives of ABH are presented and provide a proof of concept for this approach in the enhancement of enzyme-inhibitor affinity.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania , Philadelphia, Pennsylvania 19104-6323, United States.
Organizational Affiliation: