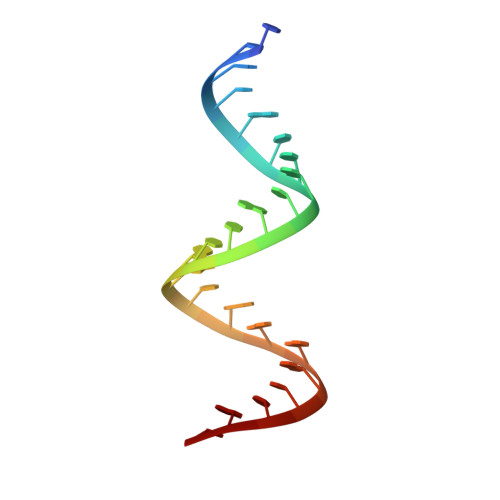
Synthesis, broad spectrum antibacterial activity, and X-ray co-crystal structure of the decoding bacterial ribosomal A-site with 4'-deoxy-4'-fluoro neomycin analogs
Hanessian, S., Saavedra, O.M., Vilchis-Reyes, M.A., Maianti, J.P., Kanazawa, H., Dozzo, P., Matias, R.D., Serio, A., Kondo, J.(2014) Chem Sci 5: 4621-4632