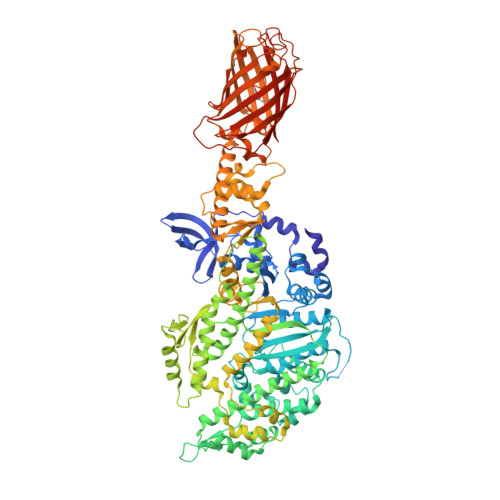
Structural basis for drug-induced allosteric changes to human beta-cardiac myosin motor activity.
Winkelmann, D.A., Forgacs, E., Miller, M.T., Stock, A.M.(2015) Nat Commun 6: 7974-7974
- PubMed: 26246073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8974
- Primary Citation Related Structures:
4PA0 - PubMed Abstract:
Omecamtiv Mecarbil (OM) is a small molecule allosteric effector of cardiac myosin that is in clinical trials for treatment of systolic heart failure. A detailed kinetic analysis of cardiac myosin has shown that the drug accelerates phosphate release by shifting the equilibrium of the hydrolysis step towards products, leading to a faster transition from weak to strong actin-bound states. The structure of the human β-cardiac motor domain (cMD) with OM bound reveals a single OM-binding site nestled in a narrow cleft separating two domains of the human cMD where it interacts with the key residues that couple lever arm movement to the nucleotide state. In addition, OM induces allosteric changes in three strands of the β-sheet that provides the communication link between the actin-binding interface and the nucleotide pocket. The OM-binding interactions and allosteric changes form the structural basis for the kinetic and mechanical tuning of cardiac myosin.
- Department of Pathology and Laboratory Medicine, Robert Wood Johnson Medical School, Rutgers University, Piscataway, New Jersey 08854, USA.
Organizational Affiliation: