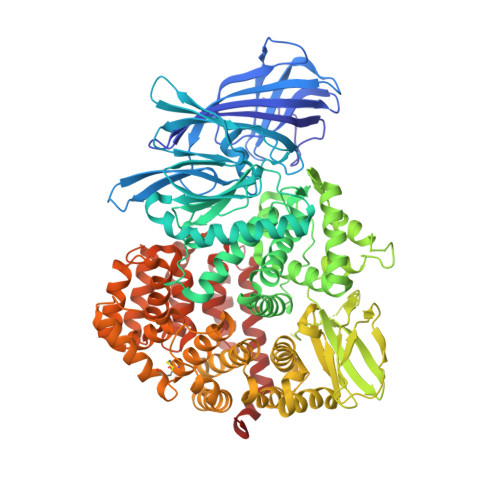
Crystal structure of human insulin-regulated aminopeptidase with specificity for cyclic peptides.
Hermans, S.J., Ascher, D.B., Hancock, N.C., Holien, J.K., Michell, B.J., Yeen Chai, S., Morton, C.J., Parker, M.W.(2015) Protein Sci 24: 190-199
- PubMed: 25408552 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2604
- Primary Citation Related Structures:
4P8Q, 4PJ6 - PubMed Abstract:
Insulin-regulated aminopeptidase (IRAP or oxytocinase) is a membrane-bound zinc-metallopeptidase that cleaves neuroactive peptides in the brain and produces memory enhancing effects when inhibited. We have determined the crystal structure of human IRAP revealing a closed, four domain arrangement with a large, mostly buried cavity abutting the active site. The structure reveals that the GAMEN exopeptidase loop adopts a very different conformation from other aminopeptidases, thus explaining IRAP's unique specificity for cyclic peptides such as oxytocin and vasopressin. Computational docking of a series of IRAP-specific cognitive enhancers into the crystal structure provides a molecular basis for their structure-activity relationships and demonstrates that the structure will be a powerful tool in the development of new classes of cognitive enhancers for treating a variety of memory disorders such as Alzheimer's disease.
- ACRF Rational Drug Discovery Centre, St. Vincent's Institute of Medical Research, Fitzroy, Melbourne, Victoria, 3065, Australia.
Organizational Affiliation: