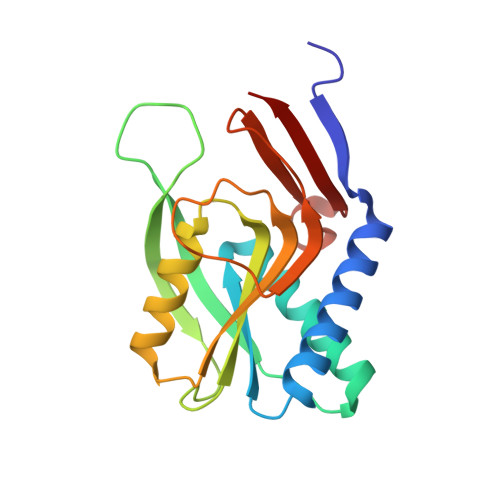
Evolution of oligomeric state through allosteric pathways that mimic ligand binding.
Perica, T., Kondo, Y., Tiwari, S.P., McLaughlin, S.H., Kemplen, K.R., Zhang, X., Steward, A., Reuter, N., Clarke, J., Teichmann, S.A.(2014) Science 346: 1254346-1254346
- PubMed: 25525255 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1254346
- Primary Citation Related Structures:
4P3K, 4P80, 4P81, 4P82, 4P83, 4P84, 4P86 - PubMed Abstract:
Evolution and design of protein complexes are almost always viewed through the lens of amino acid mutations at protein interfaces. We showed previously that residues not involved in the physical interaction between proteins make important contributions to oligomerization by acting indirectly or allosterically. In this work, we sought to investigate the mechanism by which allosteric mutations act, using the example of the PyrR family of pyrimidine operon attenuators. In this family, a perfectly sequence-conserved helix that forms a tetrameric interface is exposed as solvent-accessible surface in dimeric orthologs. This means that mutations must be acting from a distance to destabilize the interface. We identified 11 key mutations controlling oligomeric state, all distant from the interfaces and outside ligand-binding pockets. Finally, we show that the key mutations introduce conformational changes equivalent to the conformational shift between the free versus nucleotide-bound conformations of the proteins.
- European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, UK. Medical Research Council (MRC) Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge Biomedical Campus, Cambridge CB2 0QH, UK.
Organizational Affiliation: