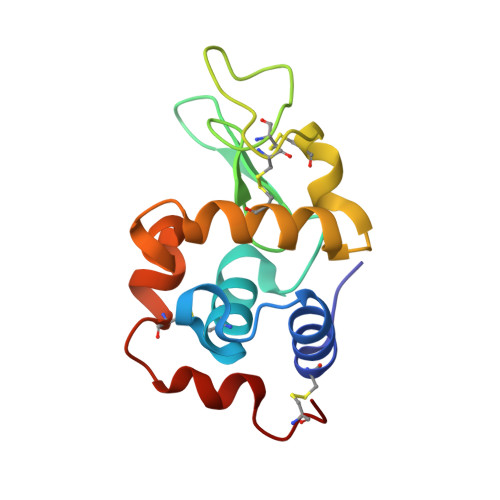
The binding of platinum hexahalides (Cl, Br and I) to hen egg-white lysozyme and the chemical transformation of the PtI6 octahedral complex to a PtI3 moiety bound to His15.
Tanley, S.W., Starkey, L.V., Lamplough, L., Kaenket, S., Helliwell, J.R.(2014) Acta Crystallogr F Struct Biol Commun 70: 1132-1134
- PubMed: 25195880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14014009
- Primary Citation Related Structures:
4OWC, 4OWE, 4OWH - PubMed Abstract:
This study examines the binding and chemical stability of the platinum hexahalides K2PtCl6, K2PtBr6 and K2PtI6 when soaked into pre-grown hen egg-white lysozyme (HEWL) crystals as the protein host. Direct comparison of the iodo complex with the chloro and bromo complexes shows that the iodo complex is partly chemically transformed to a square-planar PtI3 complex bound to the N(δ) atom of His15, a chemical behaviour that is not exhibited by the chloro or bromo complexes. Each complex does, however, bind to HEWL in its octahedral form either at one site (PtI6) or at two sites (PtBr6 and PtCl6). As heavy-atom derivatives of a protein, the octahedral shape of the hexahalides could be helpful in cases of difficult-to-interpret electron-density maps as they would be recognisable 'objects'.
- School of Chemistry, Faculty of Engineering and Physical Sciences, University of Manchester, Brunswick Street, Manchester M13 9PL, England.
Organizational Affiliation: