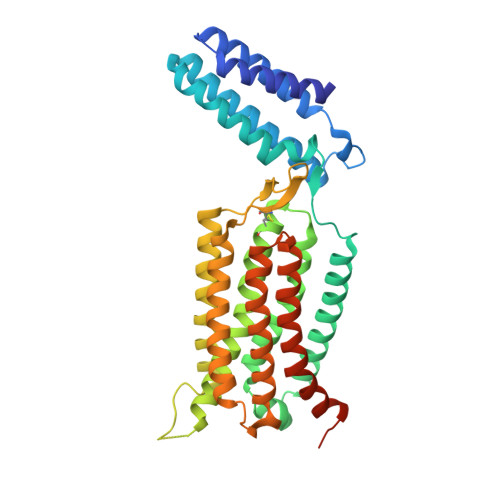
Structure of a class C GPCR metabotropic glutamate receptor 1 bound to an allosteric modulator
Wu, H., Wang, C., Gregory, K.J., Han, G.W., Cho, H.P., Xia, Y., Niswender, C.M., Katritch, V., Meiler, J., Cherezov, V., Conn, P.J., Stevens, R.C.(2014) Science 344: 58-64
- PubMed: 24603153 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1249489
- Primary Citation Related Structures:
4OR2 - PubMed Abstract:
The excitatory neurotransmitter glutamate induces modulatory actions via the metabotropic glutamate receptors (mGlus), which are class C G protein-coupled receptors (GPCRs). We determined the structure of the human mGlu1 receptor seven-transmembrane (7TM) domain bound to a negative allosteric modulator, FITM, at a resolution of 2.8 angstroms. The modulator binding site partially overlaps with the orthosteric binding sites of class A GPCRs but is more restricted than most other GPCRs. We observed a parallel 7TM dimer mediated by cholesterols, which suggests that signaling initiated by glutamate's interaction with the extracellular domain might be mediated via 7TM interactions within the full-length receptor dimer. A combination of crystallography, structure-activity relationships, mutagenesis, and full-length dimer modeling provides insights about the allosteric modulation and activation mechanism of class C GPCRs.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: