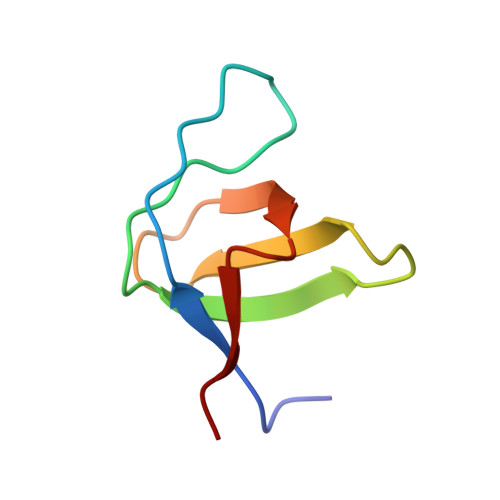
Electrostatic Effects in the Folding of the SH3 Domain of the c-Src Tyrosine Kinase: pH-Dependence in 3D-Domain Swapping and Amyloid Formation.
Bacarizo, J., Martinez-Rodriguez, S., Martin-Garcia, J.M., Andujar-Sanchez, M., Ortiz-Salmeron, E., Neira, J.L., Camara-Artigas, A.(2014) PLoS One 9: e113224-e113224
- PubMed: 25490095 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0113224
- Primary Citation Related Structures:
4JZ3, 4JZ4, 4OML, 4OMN, 4OMO, 4OMP - PubMed Abstract:
The SH3 domain of the c-Src tyrosine kinase (c-Src-SH3) aggregates to form intertwined dimers and amyloid fibrils at mild acid pHs. In this work, we show that a single mutation of residue Gln128 of this SH3 domain has a significant effect on: (i) its thermal stability; and (ii) its propensity to form amyloid fibrils. The Gln128Glu mutant forms amyloid fibrils at neutral pH but not at mild acid pH, while Gln128Lys and Gln128Arg mutants do not form these aggregates under any of the conditions assayed. We have also solved the crystallographic structures of the wild-type (WT) and Gln128Glu, Gln128Lys and Gln128Arg mutants from crystals obtained at different pHs. At pH 5.0, crystals belong to the hexagonal space group P6₅22 and the asymmetric unit is formed by one chain of the protomer of the c-Src-SH3 domain in an open conformation. At pH 7.0, crystals belong to the orthorhombic space group P2₁2₁2₁, with two molecules at the asymmetric unit showing the characteristic fold of the SH3 domain. Analysis of these crystallographic structures shows that the residue at position 128 is connected to Glu106 at the diverging β-turn through a cluster of water molecules. Changes in this hydrogen-bond network lead to the displacement of the c-Src-SH3 distal loop, resulting also in conformational changes of Leu100 that might be related to the binding of proline rich motifs. Our findings show that electrostatic interactions and solvation of residues close to the folding nucleation site of the c-Src-SH3 domain might play an important role during the folding reaction and the amyloid fibril formation.
- Department of Chemistry and Physics, Research Centre for Agricultural and Food Biotechnology (BITAL), University of Almería, Agrifood Campus of International Excellence (ceiA3), 04120, Almería, Spain.
Organizational Affiliation: