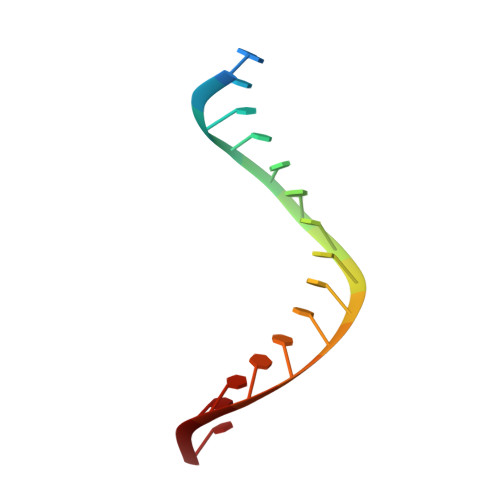
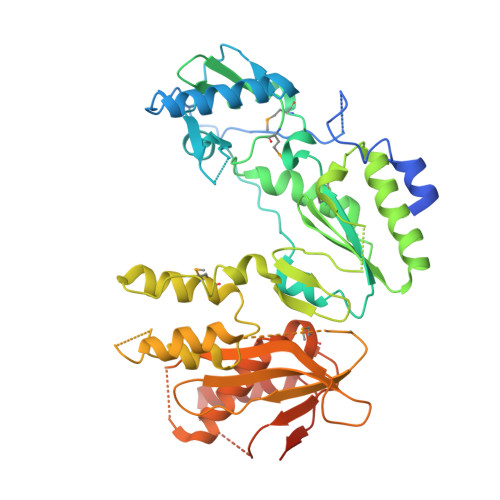
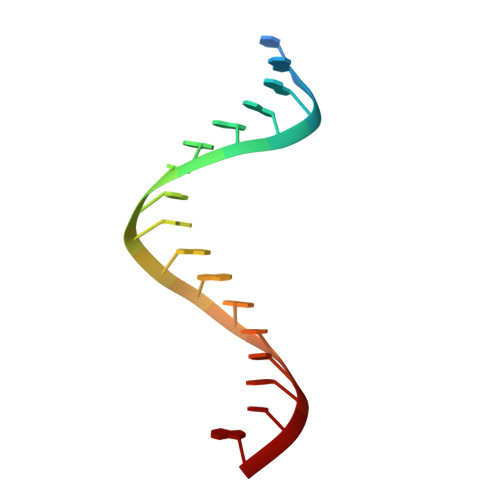
Ty3 reverse transcriptase complexed with an RNA-DNA hybrid shows structural and functional asymmetry.
Nowak, E., Miller, J.T., Bona, M.K., Studnicka, J., Szczepanowski, R.H., Jurkowski, J., Le Grice, S.F., Nowotny, M.(2014) Nat Struct Mol Biol 21: 389-396
- PubMed: 24608367 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2785
- Primary Citation Related Structures:
4OL8 - PubMed Abstract:
Retrotransposons are a class of mobile genetic elements that replicate by converting their single-stranded RNA intermediate to double-stranded DNA through the combined DNA polymerase and ribonuclease H (RNase H) activities of the element-encoded reverse transcriptase (RT). Although a wealth of structural information is available for lentiviral and gammaretroviral RTs, equivalent studies on counterpart enzymes of long terminal repeat (LTR)-containing retrotransposons, from which they are evolutionarily derived, is lacking. In this study, we report the first crystal structure of a complex of RT from the Saccharomyces cerevisiae LTR retrotransposon Ty3 in the presence of its polypurine tract-containing RNA-DNA hybrid. In contrast to its retroviral counterparts, Ty3 RT adopts an asymmetric homodimeric architecture whose assembly is substrate dependent. Moreover, our structure and biochemical data suggest that the RNase H and DNA polymerase activities are contributed by individual subunits of the homodimer.
- Laboratory of Protein Structure, International Institute of Molecular and Cell Biology, Warsaw, Poland.
Organizational Affiliation: