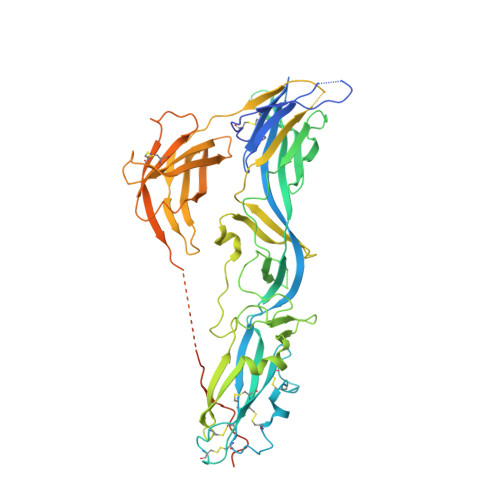
Structural basis of eukaryotic cell-cell fusion
Perez-Vargas, J., Krey, T., Valansi, C., Avinoam, O., Haouz, A., Jamin, M., Raveh-Barak, H., Podbilewicz, B., Rey, F.A.(2014) Cell 157: 407-419
- PubMed: 24725407 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2014.02.020
- Primary Citation Related Structures:
4OJC, 4OJD, 4OJE - PubMed Abstract:
Cell-cell fusion proteins are essential in development. Here we show that the C. elegans cell-cell fusion protein EFF-1 is structurally homologous to viral class II fusion proteins. The 2.6 Å crystal structure of the EFF-1 trimer displays the same 3D fold and quaternary conformation of postfusion class II viral fusion proteins, although it lacks a nonpolar "fusion loop," indicating that it does not insert into the target membrane. EFF-1 was previously shown to be required in both cells for fusion, and we show that blocking EFF-1 trimerization blocks the fusion reaction. Together, these data suggest that whereas membrane fusion driven by viral proteins entails leveraging of a nonpolar loop, EFF-1-driven fusion of cells entails trans-trimerization such that transmembrane segments anchored in the two opposing membranes are brought into contact at the tip of the EFF-1 trimer to then, analogous to SNARE-mediated vesicle fusion, zip the two membranes into one.
- Institut Pasteur, Unité de Virologie Structurale, 25-28 Rue du Docteur Roux, 75724 Paris Cedex 15, France; CNRS UMR 3569, 25-28 Rue du Docteur Roux, 75724 Paris Cedex 15, France.
Organizational Affiliation: