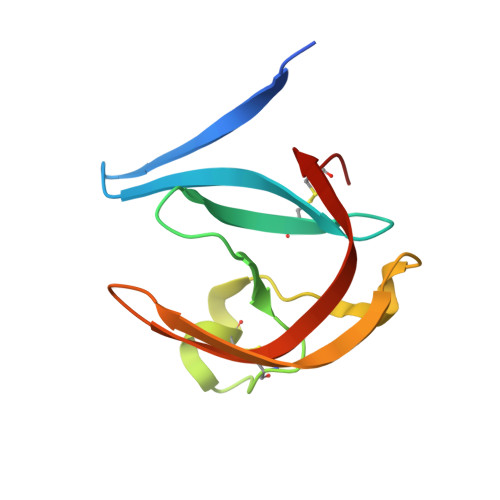
Structure-based optimization of non-peptidic Cathepsin D inhibitors.
Gradler, U., Czodrowski, P., Tsaklakidis, C., Klein, M., Werkmann, D., Lindemann, S., Maskos, K., Leuthner, B.(2014) Bioorg Med Chem Lett 24: 4141-4150
- PubMed: 25086681 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2014.07.054
- Primary Citation Related Structures:
4OBZ, 4OC6, 4OD9 - PubMed Abstract:
We discovered a novel series of non-peptidic acylguanidine inhibitors of Cathepsin D as target for osteoarthritis. The initial HTS-hits were optimized by structure-based design using CatD X-ray structures resulting in single digit nanomolar potency in the biochemical CatD assay. However, the most potent analogues showed only micromolar activities in an ex vivo glycosaminoglycan (GAG) release assay in bovine cartilage together with low cellular permeability and suboptimal microsomal stability. This new scaffold can serve as a starting point for further optimization towards in vivo efficacy.
- Merck KGaA, Merck Serono Research, Small Molecule Platform, Frankfurter Str. 250, 64293 Darmstadt, Germany. Electronic address: ulrich.graedler@merckgroup.com.
Organizational Affiliation: