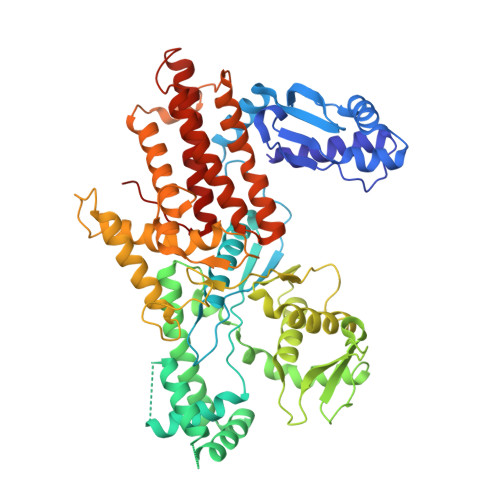
Crystal structure of E. coli arginyl-tRNA synthetase and ligand binding studies revealed key residues in arginine recognition.
Bi, K., Zheng, Y., Gao, F., Dong, J., Wang, J., Wang, Y., Gong, W.(2014) Protein Cell 5: 151-159
- PubMed: 24474195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-013-0012-1
- Primary Citation Related Structures:
4OBY - PubMed Abstract:
The arginyl-tRNA synthetase (ArgRS) catalyzes the esterification reaction between L-arginine and its cognate tRNA(Arg). Previously reported structures of ArgRS shed considerable light on the tRNA recognition mechanism, while the aspect of amino acid binding in ArgRS remains largely unexplored. Here we report the first crystal structure of E. coli ArgRS (eArgRS) complexed with L-arginine, and a series of mutational studies using isothermal titration calorimetry (ITC). Combined with previously reported work on ArgRS, our results elucidated the structural and functional roles of a series of important residues in the active site, which furthered our understanding of this unique enzyme.
- Laboratory of Non-coding RNA, Institute of Biophysics, Chinese Academy of Sciences, Beijing, 100101, China.
Organizational Affiliation: