Crystal structure of the complex between SAP97 PDZ2 and 5HT2A receptor peptide
Pandalaneni, S., Dorr, L., Mayans, O., Lian, L.-Y.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Disks large homolog 1 | 92 | Mus musculus | Mutation(s): 0 Gene Names: Dlg1, Dlgh1 | 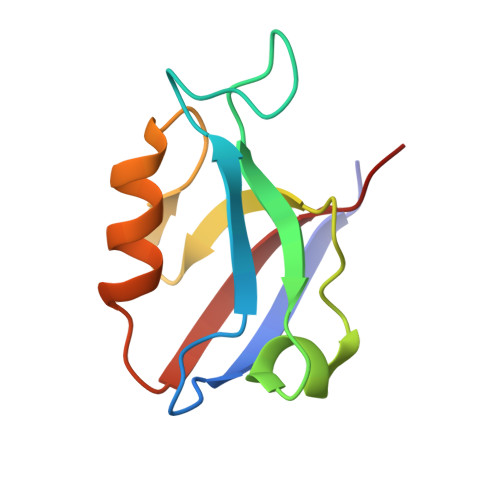 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q811D0 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 5-hydroxytryptamine receptor 2A peptide | 7 | Rattus norvegicus | Mutation(s): 0 | 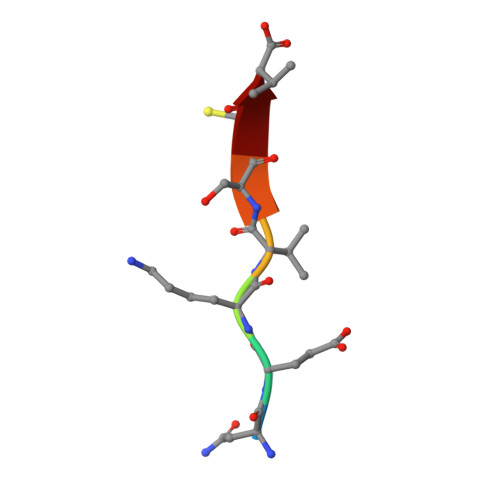 | |
UniProt | |||||
Entity Groups | |||||
| UniProt Group | P14842 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 45 | α = 90 |
| b = 45 | β = 90 |
| c = 88.11 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHASER | phasing |
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |